Open Journal of Biological Sciences
GAGA Factor Expedites Development in Drosophila
Department of Biology, Faculty of Sciences, University of Mohaghegh Ardabili, Ardabil, Iran
Author and article information
Cite this as
Das P, Bhadra MP, Bhadra U (2017) GAGA Factor Expedites Development in Drosophila. Open J Biol Sci. 2017; 2(1): 4-11. Available from: 10.17352/ojbs.000009
Copyright License
© 2017 Das P, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.The development in Drosophila is a concerted mechanism, occurring via the interplay of a constellation of genes and factors, operating in intricate synchrony. These factors, produced at precise points in their developmental cycle, operate via the activation, through binding to the various transcription factors. The GAGA factor (GAF) is such a product of the trithorax-like gene, Trl, which binds to a consensus DNA sequence for the modulation of the homeotic gene functions. Besides this, the factor has a role in chromatin remodelling; through binding with the Polycomb responsive element (PRE). The protein has a unique structural conformation with a zinc-fi nger DNA-binding, a BTB/POZ and a polyglutamine-rich Q domain. It has a unique role of acting as an anti-repressor of the gene, Kruppel, releasing the repression on it by the other DNA binding proteins. This report accomodates the interplay in which the GAGA factor is involved in the Drosophila embryonic development.
The GAGA factor (GAF), produced by trithorax-like gene, Trl, in Drosophila [1] operates by binding to its DNA recognition sequence, having a consensus sequence, GAGAG [2]. As a transcription factor, it has been found in the promoters and enhancers for the modulation of expression of various genes like the homeotic genes and developmental genes, like the Ultrabithorax (Ubx), engrailed (en), fushi-tarazu (ftz) even-skipped (eve), hsp70, hsp 26, H3/H4, Adh, E74, actin-5C, and 1-tubulin [1,3-7]. Besides these roles, the factor is also involved in remodelling of the chromatin, in functions pertaining to the Polycomb responsive element (PRE), and insulator or boundary elements [5,9–14]. This factor shows a high proportion of pleiotropism owing to its localization on the promoter regions of a multitude of genes [8,9]. The gene, Trl is presumptively produced maternally [10] hence, a better understanding of its target genes is made by studying the phenotypes using combination of hypomorphic and hypomorphic or null Trl alleles. The expression of Trl is fundamental to all the stages and all tissues of the fly, even if the levels of mRNA change significantly in these stages [11]. Till date, the studies elucidating the roles of GAGA factor have been made in the embryonic stages. Studies on larval and imago stages have been very scantily made by knocking down or over-expressing different genes. Such experiments have clearly pointed that GAGA factor is an active player in the wing disc and salivary gland development. Immuno-fluorescence studies of the salivary glands polytene chromosomes in Drosophila showed that this factor binds to a large number of euchromatic genes; hence it points out a role of GAGA factor in the maintenance of open, transcriptionally active chromatin regions.
The GAGA factor was for the first time identified [3], as an in vitro activator of the promoter of the gene, Ultrabithorax in Drosophila. Later on, it found to bind to GAGA elements (having stretches of GAGA or CTCT) over the hsp70 heat shock promoter and H3/H4 histone gene promoter regions [3,12-14]. Still further reports showed it to have crucial roles in the activation of genes and in regulating the chromatin structure [3,10,14-16]. The GAGA factor also show an interaction with the promoter sequence upstream of genes, E74 [17], his3, hid,hsp26, and hsp70 [12] in Drosophila. However, the activation of transcription is found only for the genes, Kr and Ubx. Exceptionally, Kr gene is actively transcribed when the GAGA factor binds to the anchoring site located downstream to the target Kr gene.
Structure of the GAGA protein
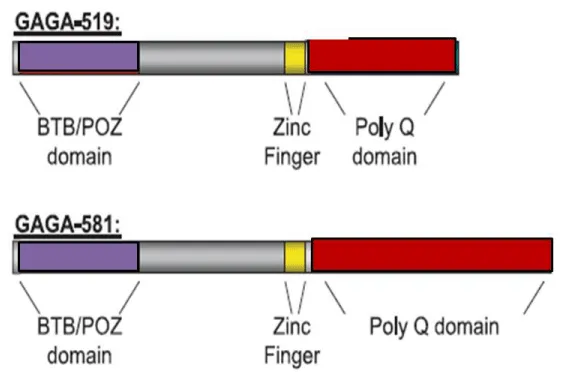
GAF bears a very unique structure having a single zinc finger DNA-binding domain, a BTB/POZ domain, a known protein–protein interaction motif [18], and a polyglutamine-rich Q domain [16,19]. The gene encodes 2 alternatively spliced isoforms of the GAF-GAGA-519 and GAGA-581 of varying length and glutamine-rich C-terminal domain sequences. Each of the isoforms have 3 distinct structural domains: a zinc finger DNA-binding domain (DBD), a broad complex tramtrack bric-a-brac/ poxvirus domain (BTB/POZ), that involves in protein–protein interaction, and the C-terminal Q domain of a glutamine-rich C-terminus area involved in transcription activation [20-22] (Figure 1). Contrary to initial notion, of the Q domain acting as a transcriptional activator [18,23,24], GAF, in fact operates as an anti-repressor. It is constituted of the following components:
The DNA-binding domain or the DBD: The C-terminus of the GAF possesses a single classical C2-H2 zinc finger DNA-binding domain of 82 amino acid residues from 310–391 after a basic helix flanked of 3 short tracts of basic amino acid residues: BR1, BR2, and BR3 [20,21]. The consensus binding sequence bears a minimal penta-nucleotide, GAGAG [21] of which the tri-nucleotide sequence GAG is essential for recognition and binding [25]. Once that a zinc finger binds the major groove, identifying the GAG sequence, it gets stabilized through interactions with an extension of basic amino acids, at the N-terminal [20,21]. The tract BR1 also wraps around the minor groove with Adenine in the fourth position of the sequence GAGAG [20].
BTB/POZ domain: The BTB/POZ domain of the GAF occurs in all groups of organisms has 122 amino acids located in the N-terminal region of the GAF. This domain provides an interface for the occurrence of protein–protein interactions during transcriptional activation and transcriptional factor repression [18,26-32]. The protein oligomers are actually the result of 3 specific residues- D35, G93, and L112 and there are 3 -helices adjacent to these residues that help in capping, stabilizing the dimerizing at interface [33]. The GAF BTB/POZ domain in GAF forms higher order functional multimeric complexes by virtue of self-association [22,33,34].
Q domain: The C-terminal bears a Q domain that have been involved in promoter distortion, single-strand binding and multi-merization [35]. Because of the single-stranded DNA binding affinity, GAF is able to interact with triple-stranded DNA [36]. In Drosophila S2 cells, the Q domain was found to play a role of trans-activation domain. Combinatorial studies indicate that the Q domain is primarily involved in the larger GAF complex formation, but is not needed for heat shock response functions of the gene, hsp83 [19,35].
DNA–binding activity of the GAGA factor
When the GAGA factor binds to the major groove of DNA and the zinc finger of DBD, additionally interacting with the basic region BR1 at the minor groove [20], it results in a distorted DNA via the interactions of the Q domain at the promoter regions that melts the duplexes [35], especially when GAF binds to the single- and triple-stranded DNA over stretches of (GA)n [35,36]. The DNA binding over multiple GA or CA stretches is due to the multimer formation caused by Q domain [35] and the BTB/POZ domain [34]. GAF binding sites are more frequent in the introns that suggest a possibility of regulatory role in transcriptional elongation [37]. The protein may to bind to long GA-rich repeats as evident in case of satellite DNA [38,39]. GAF can cause the maintenance of heterochromatic regions in the transcriptionally repressed state by recruiting arrays of GAF at the high density binding sites for GAF.
Interacting partners of GAGA factor
Being a multi-functional protein, GAF, can interact with a significant number of partners executing functions apart from binding single, double, and triple stranded DNA, such as providing multiple binding sites and support for the cofactors to form a feasible functional complex. GAF has depicted to possess the ability to show self-associate as well as associate with other BTB/POZ functional proteins like tramtrack (Ttk), Pipsqueak (Psq), and batman (ban) [6,40,41].
Tramtrack
Tramtrack or ttk, is a gene regulating development, that codes for a protein important in oocyte development [A factor that regulates the class II major histocompatibility complex gene DPA is a member of a subfamily of zinc finger proteins that includes a Drosophila developmental control protein [42]. The protein, Ttk may be expressed as two proteins-p69 and p88, that bind to the regulatory regions of several segmentation genes [42], like ftz to show novel expression pattern in embryos. During the transition from the mitotic to endocycle in the follicle cells, ttk promotes the activity of JNK pathway by passing the notch signalling pathway [43]. The ttk also codes for proteins that represses the expression of engrailed with the help of runt [44]. The ttk, thus represses several Drosophila genes, the activity of such genes are counteracted by GAF in vivo, which functions as anti-repressor [6].
Pipsqueak (Psq)
Psq, essentially regulates development via recognition of the GAGA sequences. However, it needs the GA stretch to be longer than the GAF [2]. Psq has been found to co-localize with GAF for numerous loci in the polytene chromosomes [40], where it and GAF, being members of the polycomb group of complexes, bind to the polycomb response element of the bxd gene [45,46]. The interaction between them occurs through their BTB/POZ domains [40].
Batman or ban
The batman or ban co-localizes with GAF and causes the activation and repression of homeotic genes [41]. Besides ban, the proteins, Corto and the sin3-associated polypeptide SAP18, form complexes with GAF’s BTB/POZ domain [47,48] and causes histone deacetylation. GAF interacts with the large subunit, NURF 301 of the NURF [49,50], dSSRP1 subunit of FACT [51,52].
GAGA arrests the spreading of heterochromatin
Developmental activities of an organism are under the stringent epigenetic maintenance of the genomic expression. The complex called “facilitates chromatin transcription” or FACT in Drosophila, interact with GAGA factor to modulate the structure of the chromatin expression. The gene expression is epigenetically maintained through post-translational modifications of the four core histones on the nucleosome [53]. The expression of two such epigenetic genes- the Hox gene, whose expression is governed by Polycomb and trithorax group genes [54,55] and the position effect variegation (PEV) [56,57], has been extensively studied. Both these phenomena involve silencing and counteracting the maintenance of the active state. The methylation of histone H3 at K27 and/or K9 cause silencing, which then leads to the binding of the Polycomb group or heterochromatin proteins that recognize these silent marks [55,57-60].
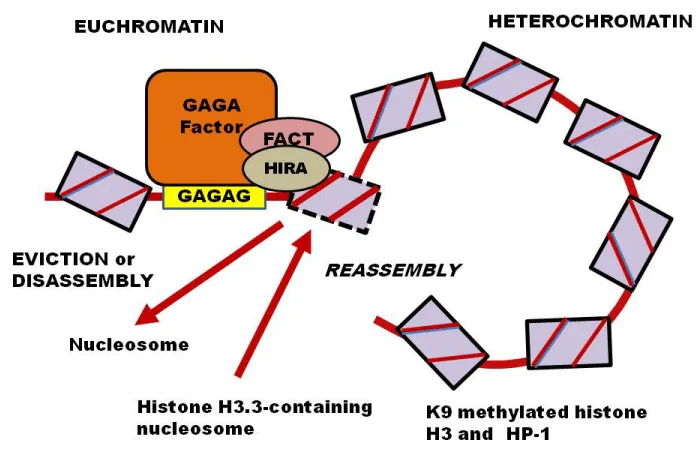
The GAF and FACT replace histone H3 into H3.3, by associating HIRA with d1, resulting in the maintenance of expression of the white gene in the heterochromatic environment. The active state is sustained through the replacement of Histone H3.3 by the removal of a nucleosome, replacing with H3.3 containing nucleosome with stepwise assembling and disassembling of a nucleosome at the DNase-hypersensitive site of d1. Heterochromatin formation is marked by K9-methylated histone H3 and its binding protein HP1, and has a tendency to spread into neighbouring regions [57,58]. The process of reassembling of the nucleosome, following the replacement of histone, keep on removing K9-methylated histone H3 at d1 and prevent the spreading of the heterochromatin (Figure 2).
GAGA in primordial germ cell migration and gonad development
The development of gonad is possible through the migration of primordial germ cells (PGCs). The germ cell progenitors are formed at the posterior pole of the embryo in the region where the maternal germplasm reside. Once specified, after several mitotic divisions, these cells separate from the embryonic syncitium and move inside the embryo [61,62]. The migration of PGC has two stages- passive and active. While passive migration occurs during morphogenetic events of gastrulation, leading to the shifting of germ cells to the midgut pocket by cellular invagination of the prospective endoderm, active migration takes place within the embryo via the autonomous movement of the PGCs. PGCs cross the primary gut wall forming pseudopodia like structures and then join the mesodermal cells to form the somatic progenitor cells of the gonads, localized to either sides of the ventral groove in the fifth abdominal embryonic segment [61-63].
The GAGA factor encoding gene, Trl is expressed in various stages of Drosophila are found to regulate several developmental processes like embryogenesis, oogenesis, eye and wing development, and formation of dorsal processes [2,10,64-67]. In developing male germ cells in Drosophila, a loss-of-function in Trl has been found to lead to partial germ cell loss [68]. A recent study pointed that a mutant for Trl shows an early activation of migration and movement to the inward of the embryo. Such a premature drift lead to loss of orientation and an absence of a normal gonad formation. The migrations of the cells are under the JAK-STAT signalling pathway factors that activate the GPCR coded by gene tre1 or trapped in endoderm 1 [69-71], whose product is normally limited to the germ cells. In some cases, the somatic environment can modulate spatiotemporal regulation of germ cell migration as in mutants of the hopscotch gene, encoding a Janus kinase, causing a premature migration of primordial cells [72]. This suggests that the GAGA factor does influence the migration of primordial cells via their contact with the somatic cells.
GAGA in wing development
The over-expression of GAF has been found to alter the gene expression of many genes in the wing disc. Depletion in GAGA, caused by the deletion at 69B was found to consistently reduce the size of the wing by about 10 %. Using NubbGAL4 with Dicer2 reduction was by 55% and in case of ptcGAL4 by 35%. Presumptively the wing size diminished was due to the defect in the proliferation of the cell. The abdominal segment A6 showed a transformation to A5, in males having a depleted GAGA factor. It is well known fact that a defective Abd-B is responsible for homeotic transformation of the abdominal segments. The above findings thus correlate that GAGA factor was capable of initiating a defective transformation of segmentation of the embryo. A heterozygous null Trl male was found to have a similar transformation owing to the deficiency (Trl67/Df(3R)Sbd26) [48]. A phenotype of loss-of-function due to usage of pannier GAL4 (pnrGAL4) showed a cleft with loss of bristle at the dorsal of the notum while over-expression was found to completely affect the dorsal closure with embryonic death before the first instar stage is reached.
GAGA factor as anti-repressor of Kruppel
Kruppel is the Drosophila gap gene that plays a crucial role in the early embryonic development by forming the antero-posterior boundaries. The syncitial blastoderm stage shows the expression of the Kr in the form of a circular band girdling around 45 to 55% of the egg [73]. The Kruppel transcripts are formed at around 2 to 5 hr later from hatching [74] and protein coded thereafter bears zinc fingers [75-77] that in turn regulate the expression of the other genes, both spatially and temporally.
The analysis of the proximal promoter of the Drosophila Kruppel (Kr) gene shows a 44-base pair length fragment bearing the RNA start sites having prominent promoter activity. This promoter has both upstream and downstream flanked by sites binding with the GAGA factor. The GAF interacts with the Kr promoter region downstream in a sequence-specific fashion, and the purified protein is found to activate the in vitro transcription of Kr and Ubx. The GAF is acting as an anti-repressor acting in the presence of the binding site to repress the inactivation of the Kr by undoing the repression by the DNA binding factor. The transcriptional anti-repression model of the gene describes that DNA binding factors may bind with the gene’s promoter and enhancer sequence repressing the gene function. GAGA factor is able to nullify the repressive effect of such factors resulting in allowing transcription of Kr.
GAGA factor relieves the repression of tailless
The development of both the anterior and posterior poles, which are the terminal domains of Drosophila embryos, is specified by the maternal terminal system [78]. One of such gene is tailless, which is crucial for the development of terminal structure like telson and the posterior gut as well as head portions as head structures and the brain development [79-82]. The syncitial blastoderm stage of the embryo expression this transcript at the poles occur after the indirect activation by the maternally produced Torso receptor tyrosine kinase pathway at the embryonic termini. It partly relieves the repression caused by the HMG transcription repressor, Capicua and the co-repressor, Groucho [83,84]. The other repressor that it negates is the BTB domain zinc finger protein, Tramtrack69 [85]. A successful functional tailless ensures the normal expression of of other gap genes such as Kruppel and Knirps, and later on genes like hunchback , brachyenteron, and forkhead [79-82]. The reduction of concentration of the repressor causes a loss in a well-defined edge of expression domains [86,87]. It is seen that if the binding affinity of GAGA factor to the tor-RE is low, and multiple tor-REs are present in the tll cis-regulatory region [85,88] the boundary of tll expression gets poorly-defined.
Zelda as a candidate for GAGA
The DNA-binding protein Zelda or Vielfaltig in Drosophila is an active transcriptional activator of the zygotic genome and produces an open chromatin state. Once that, the chromatin open, it facilitates the recruitment of the transcriptional factor. This leads to the remodelling of the genome, causing the target gene to express. But sometimes absence of the Zelda can also lead to transcriptional activation in case of factors like the GAGA factor binding motif and the GAGA factor in embryo. The 14th nuclear cycle is the testimony to the initiation of zygotic transcription, where Zelda is key activator of the zygotic genome in the maternal to zygotic transition [89,90]. Zelda binding sites have been found to be critical for regulation of DNA binding by transcriptional factors Dorsal (Dl), Twist (Twi), and Bicoid (Bcd) [91-93]. The Zelda also potentiates transcription factor binding sites by determining sites of open chromatin [94,95]. The density of histone H3 increases when the Zelda decreases in wild-type embryos [96], the Zelda thus, dictate the expression of the initial set of zygotic genes, transcribed post fertilization and also binds to the locus for genes that need to be activated later such that a precise sequence of gene activation ensues during gastrulation.
The Vertebrate homologue of GAGA factor
Although the essential and conserved role of PcG/trxG homolog was clearly proven in the Drosophila melanogaster, the vertebrate homologue for the Drosophila GAGA factor was unknown until the recent studies [55]. The recognition sites for the GAGA factor called the GAGA boxes were found in many genes in the vertebrates including the hox complexes but a putative GAGA factor was yet to be discovered [16,97-101].Very recently, the vertebrate GAG factor homologue having properties similar to the Drosophila in terms of domain structure and capacity for DNA recognition and binding. Various analysis including the structural modelling, phylogenetic analysis and cross-reactivity studies have exposed that GAGA factor can bind to DNA sequences rich in GAGA in the hox complex. In mouse and human, cKrox (Kruppel-related zinc finger protein cKrox) or Th-POK (T-helper inducing POZ/Kruppel-like factor) was deemed as the homolog of the Drosophila GAGA factor. It is found to be encoded by the zbtb7b or Zfp67 gene. Th-POK mainly regulates the commitment towards the lineages CD4 and CD8. It is found that the mice having a mutated ArgineY Glycine in the X position of the second zinc finger are of Th-POK show immune-compromisation.This means that the Arginine is responsible for the recognition of the invariant G of the target and any mutation in the third position of the consensus sequence, GAGAG alters the target binding specificity [20]. This highlights the role of G in binding and arginine in specific DNA binding, both in fly as well as in mammals. The binding sites of c-Krox/Th-POK are found to be rich in purine with the pentamer sequence GAGA in the target. Some studies have also shown that Th-POK/c-Krox binds to the collagen promoter region to cause transcriptional activation [102-104] and the deletion of the C-terminal region, the transcriptional activity is reduced in mice [105].
The Evx2 and Hoxd13 genes have tracts rich in GA in mouse, human, and zebrafish and functions in blocking the enhancer in both transgenic flies and cultured human cells [106]. Mutating the GAGA binding sequence prevents it from functioning as an insulator [106]. The murine Hox clusters with Histone free regions associate with the GAF recognition sites and regulate the binding of Th-POK. Thus, the mammalian GAGA factor act in nucleosome reorganization at the Hox clusters providing a platform for binding regulatory proteins, organizing chromatin regulatory activities of the chromatin, including the formation of boundaries.
Conclusion
The evidence cited in this report, clearly indicates that the GAGA factor is involved at several levels in gene expression regulation. Hence, it would be improper to consider it as a simple transcription factor or anti-repressor. Its role as a structural protein on the chromatin conformation, from its primary to its tertiary structure, depicts a potential role as a transcriptional activator and repressor. The actual role of GAF in maintaining the secondary and tertiary structure still remains more speculative than quantitative; even the functional significance of GAF multimers still remains cryptic. The multimers may affect the topology of the regulatory region where they bind, changing the rotational phase of the nucleosomes to enable the proteins to co-interact. The purification of GAF and the fully characterized chromatin system may reveal the full structural analysis data. The chromatin folding and cis-trans interaction of the regulatory sequences may also give additionally unique information about the effect of the GAGA factor in the folding of the chromatin.
The diversified role of the GAGA factor in binding sequence-specifically to chromatin, DNA, transcriptional factors, metal ions and leading to protein homo-dimerization as well as hetero-dimerization has very critical roles in the normal execution of biological processes like the cell division, chromatin assembly, chromatin modification, chromatin organization, dosage compensation, imaginal disc-derived wing morphogenesis, mitotic nuclear division, negative regulation of transcription, nuclear division, oogenesis, positive regulation of chromatin silencing, positive regulation of transcription, positive regulation of transcription from RNA polymerase II promoter, protein oligomerization, sensory perception of pain, spermatogenesis and syncytial blastoderm mitotic cell cycle. Such a multi-faceted protein does require further detailed analysis for still some functions not yet discerned completely. The numerous avenues that this factor enlightens its role in, clearly pinpoint the possibility of exploiting this factor in abating some diseases that are caused by an aberrant cell cycle progression, like the various cancers, cell proliferation, tumor development and pattern formation, during embryogenesis. Thus, it still requires focussed and multi-disciplinary efforts to dissect the yet unknown transcriptional regulatory mechanisms that regulate Drosophila development.
The work was supported by an HFSP Young Investigator Grant (RGY020), CSIR Net work Project (BSC 0108, 0121) awarded to UB and Wellcome Trust International Fellowship awarded to MPB (GAP0065).
Authors’ contributions
The initial write-up was done by PD. Figures were drawn by PD. Final write-up and the concept was contributed by UB and MPB.
- Granok H, Leibovitch BA, Shaffer CD, Elgin SC (1995) Chromatin: Ga-ga over GAGA factor. Curr Biol 5: 238-241. Link: https://goo.gl/88kMSE
- Lehmann M, Siegmund T, Lintermann KG, Korge G (1998) The Pipsqueak Protein of Drosophila melanogasterBinds to GAGA Sequences through a Novel DNA-binding Domain. J Biol Chem 273: 28504-28509. Link: https://goo.gl/wg50mU
- Biggin MD, Tjian R (1988) Transcription factors that activate the Ultrabithorax promoter in developmentally staged extracts. Cell 53: 699-711. Link: https://goo.gl/ypVMv1
- Ohtsuki S, Levine M (1998) GAGA mediates the enhancer blocking activity of the eve promoter in the Drosophila embryo. Genes Dev 12: 3325-3330. Link: https://goo.gl/UYU8B1
- Okada M, Hirose S (1998) Chromatin remodeling mediated by Drosophila GAGA factor and ISWI activates fushi tarazu gene transcription in vitro. Mol Cell Biol 18: 2455-2461. Link: https://goo.gl/NF1uql
- Pagans S, Ortiz-Lombardía M, Espinás ML, Bernués J, Azorín F (2002) The Drosophila transcription factor tramtrack (TTK) interacts with Trithorax ‐like (GAGA) and represses GAGA‐mediated activation. Nucleic Acids Res 30: 4406-4413. Link: https://goo.gl/qzW5Zk
- Read D, Nishigaki T, Manley J (1990) The Drosophila even-skipped promoter is transcribed in a stage-specific manner in vitro and contains multiple, overlapping factor-binding sites. Mol Cell Biol 10: 4334-4344. Link: https://goo.gl/x7cp9k
- Lee C, Li X, Hechmer A, Eisen M, Biggin MD, et al. (2008) NELF and GAGA factor are linked to promoter-proximal pausing at many genes in Drosophila. Mol Cell Biol 28: 3290-3300. Link: https://goo.gl/eaADYH
- Schuettengruber B, Ganapathi M, Leblanc B, Portoso M, Jaschek R, et al. (2009) Functional anatomy of polycomb and trithorax chromatin landscapes in Drosophila embryos. PLoS Biol 7: e1000013. Link: https://goo.gl/opDcGS
- Bhat KM, Farkas G, Karch F, Gyurkovics H, Gausz J, et al. (1996) The GAGA factor is required in the early Drosophila embryo not only for transcriptional regulation but also for nuclear division. Development 122: 1113-1124. Link: https://goo.gl/I9AT5a
- Graveley BR, Brooks AN, Carlson JW, Duff MO, Landolin JM, et al. (2011) The developmental transcriptome of Drosophila melanogaster. Nature 471: 473-479. Link: https://goo.gl/Iz7jyu
- Gilmour DS, Thomas GH, Elgin SC (1989) Drosophila nuclear proteins bind to regions of alternating C and T residues in gene promoters. Science 245: 1487-1490. Link: https://goo.gl/FqvLg9
- Lee H, Kraus KW, Wolfner MF, Lis JT (1992) DNA sequence requirements for generating paused polymerase at the start of hsp70. Gens Dev 6: 284-295. Link: https://goo.gl/WKQxsN
- Tsukiyama T, Becker PB, Wu C (1994) ATP-dependent nucleosome disruption at a heat-shock promoter mediated by binding of GAGA transcription factor. Nature 525-532. Link: https://goo.gl/rke34R
- Farkas G, Gausz J, Galloni M, Reuter G, Gyurkovics H, et al. (1994) The Trithorax-like gene encodes the Drosophila GAGA factor. Nature 371: 806-808 Link:
- Lehmann M (2004) Anything else but GAGA: a nonhistone protein complex reshapes chromatin structure. TRENDS in Genetics 20: 15-22. Link: https://goo.gl/UOOLQG
- Thummel CS (1989) The Drosophila E74 promoter contains essential sequences downstream from the start site of transcription. Gens Dev 3: 782-792. Link: https://goo.gl/kNM7ZV
- Bardwell VJ, Treisman R (1994) The POZ domain: a conserved protein-protein interaction motif. Gens Dev 8: 1664-1677. Link: https://goo.gl/kfdqST
- Greenberg AJ, Schedl P (2001) GAGA factor isoforms have distinct but overlapping functions in vivo. Mol Cell Biol 21: 8565-8574. Link: https://goo.gl/9fMSBI
- Omichinski JG, Pedone PV, Felsenfeld G, Gronenborn AM, Clore GM (1997) The solution structure of a specific GAGA factor-DNA complex reveals a modular binding mode. Nat Struct Biol 4: 122-132. Link: https://goo.gl/ftdixx
- Pedone PV, Ghirlando R, Clore GM, Gronenborn AM, Felsenfeld G, et al. (1996) The single Cys2-His2 zinc finger domain of the GAGA protein flanked by basic residues is sufficient for high-affinity specific DNA binding. Proc Natl Acad Sci U S A 93: 2822-2826. Link: https://goo.gl/n8lgPV
- Katsani KR, Hajibagheri MA, Verrijzer CP (1997) Co ‐operative DNA binding by GAGA transcription factor requires the conserved BTB/POZ domain and reorganizes promoter topology. The EMBO journal 18: 698-708. Link: https://goo.gl/Fz1TAH
- Miller MW (1985) Cogeneration of retrogradely labeled corticocortical projection and GABA-immunoreactive local circuit neurons in cerebral cortex. Brain Res 23: 187-192. Link: https://goo.gl/tDaz9Q
- Chen LQ, Santarelli V, Horn R, Kallen RG (1996) A unique role for the S4 segment of domain 4 in the inactivation of sodium channels. J Gen Physiol 108: 549-556. Link: https://goo.gl/C4imZo
- Wilkins RC, Lis JT (1998) GAGA factor binding to DNA via a single trinucleotide sequence element. Nucleic Acids Res 26: 2672-2678. Link: https://goo.gl/ky4nph
- DiBello PR, Withers DA, Bayer CA, Fristrom JW, Guild GM (1991) The Drosophila Broad-Complex encodes a family of related proteins containing zinc fingers. Genetics 129: 385-397. Link: https://goo.gl/ggZeQ1
- Godt D, Couderc JL, Cramton SE, Laski FA (1993) Pattern formation in the limbs of Drosophila: bric a brac is expressed in both a gradient and a wave-like pattern and is required for specification and proper segmentation of the tarsus. Development 119: 799-812. Link: https://goo.gl/vOZrJ4
- Read D, Manley JL (1992) Alternatively spliced transcripts of the Drosophila tramtrack gene encode zinc finger proteins with distinct DNA binding specificities. EMBO J 11: 1035-1044. Link: https://goo.gl/C3Ec6Q
- Deweindt C, Albagli O, Bernardin F, Dhordain P, Quief S, et al. (1995) The LAZ3/BCL6 oncogene encodes a sequence-specific transcriptional inhibitor: a novel function for the BTB/POZ domain as an autonomous repressing domain. Cell Growth Differ 6: 1495-1503. Link: https://goo.gl/ewNdnT
- Chang CC, Ye BH, Chaganti RS, Dalla-Favera R (1996) BCL-6, a POZ/zinc-finger protein, is a sequence-specific transcriptional repressor. Proc Natl Acad Sci 93: 6947-6952. Link: https://goo.gl/93b4CC
- Seyfert VL, Allman D, He Y, Staudt LM (1996) Transcriptional repression by the proto-oncogene BCL-6. Oncogene 12: 2331-2342. Link: https://goo.gl/nn9KJ1
- Kaplan J, Calame K (1997) The ZiN/POZ domain of ZF5 is required for both transcriptional activation and repression. Nucleic Acids Res 25: 1108-1116. Link: https://goo.gl/z0tKhm
- Read D, Butte MJ, Dernburg AF, Frasch M, Kornberg TB (2000) Functional studies of the BTB domain in the Drosophila GAGA and Mod (mdg4) proteins. Nucleic Acids Res 28: 3864-3870. Link: https://goo.gl/5UMLrJ
- Espinás ML, Jiménez-García E, Vaquero A, Canudas S, Bernués J, et al. (1999) The N-terminal POZ domain of GAGA mediates the formation of oligomers that bind DNA with high affinity and specificity. J Biol Chem 274: 16461-16469. Link: https://goo.gl/u9oF69
- Wilkins RC, Lis JT (1999) DNA distortion and multimerization: novel functions of the glutamine-rich domain of GAGA factor. J Molecu Biol 285: 515-525. Link: https://goo.gl/rwu9tp
- Jiménez-García E1, Vaquero A, Espinás ML, Soliva R, Orozco M, et al. (1998) The GAGA factor of Drosophila binds triple-stranded DNA. J Biol Chem 273: 24640-24648. Link: https://goo.gl/auJGr5
- van Steensel B, Delrow J, Bussemaker HJ (2003) Genomewide analysis of Drosophila GAGA factor target genes reveals context-dependent DNA binding. Proc Natl Acad Sci USA 100: 2580-2585. Link: https://goo.gl/5QHEdo
- Platero JS, Csink AK, Quintanilla A, Henikoff S (1998) Changes in chromosomal localization of heterochromatin-binding proteins during the cell cycle in Drosophila. J Cell Biol 140: 1297-1306. Link: https://goo.gl/URPcbB
- Raff JW, Kellum R, Alberts B (1994) The Drosophila GAGA transcription factor is associated with specific regions of heterochromatin throughout the cell cycle. EMBO J 13: 5977-5983. Link: https://goo.gl/aSDnsL
- Schwendemann A, Lehmann M (2002) Pipsqueak and GAGA factor act in concert as partners at homeotic and many other loci. Proc Natl Acad Sci USA 99: 12883-12888. Link: https://goo.gl/BW0YQ1
- Faucheux M, Roignant JY, Netter S, Charollais J, Antoniewski C, et al. (2003) batman Interacts with polycomb and trithorax group genes and encodes a BTB/POZ protein that is included in a complex containing GAGA factor. Mol Cell Biol 23: 1181-1195. Link: https://goo.gl/1Luk5h
- Xiong WC, Montell C (1993) tramtrack is a transcriptional repressor required for cell fate determination in the Drosophila eye. Gens Dev 7: 1085-1096. Link: https://goo.gl/izOHh5
- Jordan KC, Schaeffer V, Fischer KA, Gray EE, Ruohola-Baker H, et al. (2006) Notch signaling through tramtrack bypasses the mitosis promoting activity of the JNK pathway in the mitotic-to-endocycle transition of Drosophila follicle cells. BMC Dev Biol 6: 16. Link: https://goo.gl/jnVwhn
- Vander Zwan CJ, Wheeler JC, Li LH, Tracey WD, Gergen JP (2003) A DNA-binding-independent pathway of repression by the Drosophila Runt protein. Blood Cells Mol Dis 30: 207-222. Link: https://goo.gl/TVmRW6
- Horard B, Tatout C, Poux S, Pirrotta V (2002) Structure of a polycomb response element and in vitro binding of polycomb group complexes containing GAGA factor. Mol Cell Biol 20: 3187-3197. Link: https://goo.gl/d37sGW
- Huang DH, Chang YL, Yang CC, Pan IC, King B (2002) Pipsqueak encodes a factor essential for sequence-specific targeting of a polycomb group protein complex. Mol Cell Biol 22: 6261-6271. Link: https://goo.gl/HPlfCB
- Salvaing J, Lopez A, Boivin A, Deutsch JS, Peronnet F (2003) The Drosophila Corto protein interacts with Polycomb ‐group proteins and the GAGA factor. Nucleic Acids Res 31: 2873-2882. Link: https://goo.gl/tYfdYb
- Espinás ML, Canudas S, Fanti L, Pimpinelli S, Casanova J, et al. (2000) The GAGA factor of Drosophila interacts with SAP18, a Sin3 ‐associated polypeptide. EMBO Rep 1: 253-259. Link: https://goo.gl/OH7xdy
- Xiao H, Sandaltzopoulos R, Wang HM, Hamiche A, Ranallo R, et al. (2001) Dual functions of largest NURF subunit NURF301 in nucleosome sliding and transcription factor interactions. Mol Cell 8: 531-543. Link: https://goo.gl/ynrnUu
- Badenhorst P, Voas M, Rebay I, Wu C (2002) Biological functions of the ISWI chromatin remodeling complex NURF. Gens Dev 16: 3186-3198. Link: https://goo.gl/EtJ5mv
- Orphanides G, LeRoy G, Chang CH, Luse DS, Reinberg D (1998) FACT, a factor that facilitates transcript elongation through nucleosomes. Cell 92: 105-116. Link: https://goo.gl/ymfSY4
- Shimojima T, Okada M, Nakayama T, Ueda H, Okawa K, et al. (2003) Drosophila FACT contributes to Hox gene expression through physical and functional interactions with GAGA factor. Gens Dev 17: 1605-1616. Link: https://goo.gl/QZBawn
- Jenuwein T, Allis CD (2001) Translating the histone code. Science 293: 1074-1080. Link: https://goo.gl/cnDmgP
- Simon JA, Tamkun JW (2002) Programming off and on states in chromatin: mechanisms of Polycomb and trithorax group complexes. Curr Opin Genet Dev 12: 210-218. Link: https://goo.gl/PHZ92K
- Ringrose L, Paro R (2004) Epigenetic regulation of cellular memory by the Polycomb and Trithorax group proteins. Annu Rev Genet 38: 413-443. Link: https://goo.gl/MVHYJd
- Reuter G1, Spierer P (1992) Position effect variegation and chromatin proteins. Bioessays 14: 605-612. Link: https://goo.gl/TCYddA
- Grewal SI, Elgin SC (2002) Heterochromatin: new possibilities for the inheritance of structure. Curr Opin Genet Dev 12: 178-187. Link: https://goo.gl/wPXcPD
- Rea S, Eisenhaber F, O'Carroll D, Strahl BD, Sun ZW, et al. (2000) Regulation of chromatin structure by site-specific histone H3 methyltransferases. Nature 406: 593-599. Link: https://goo.gl/LbbJf2
- Müller J, Hart CM, Francis NJ, Vargas ML, Sengupta A, et al. (2002) Histone methyltransferase activity of a Drosophila Polycomb group repressor complex. Cell 111: 197-208. Link: https://goo.gl/Z0UP93
- Czermin B, Melfi R, McCabe D, Seitz V, Imhof A, et al. (2002) Drosophila enhancer of Zeste/ESC complexes have a histone H3 methyltransferase activity that marks chromosomal Polycomb sites. Cell 111: 185-196. Link: https://goo.gl/W6Diy4
- Dansereau DA, Lasko P (2008) The development of germline stem cells in Drosophila. Germline Stem Cells 3-26. Link: https://goo.gl/ppS1Cb
- Richardson BE, Lehmann R (2010) Mechanisms guiding primordial germ cell migration: strategies from different organisms. Nat Rev Mol Cell Biol 11: 37-49. Link: https://goo.gl/ndj4I3
- Jemc JC (2011) Somatic gonadal cells: the supporting cast for the germline. Genesis 49: 753-775. Link: https://goo.gl/NUShM5
- Ogienko AA, Karagodin DA, Pavlova NA, Fedorova SA, Voloshina MV, et al. (2008) Molecular and genetic description of a new hypomorphic mutation of Trithorax-like gene and analysis of its effect on Drosophila melanogaster oogenesis. Russian journal of developmental biology 39: 108-115. Link: https://goo.gl/rtCId1
- Ogienko AG, Kurnosov AV, Manakov AY, Larionov EG, Ancharov AI, et al. (2006) Gas hydrates of argon and methane synthesized at high pressures: composition, thermal expansion, and self-preservation. J Phys Chem B 110: 2840-2846. Link: https://goo.gl/oWQY3n
- Omelina ES, Baricheva EM, Oshchepkov DY, Merkulova TI, et al. (2011) Analysis and recognition of the GAGA transcription factor binding sites in Drosophila genes. Comput Biol Chem 35: 363-370. Link: https://goo.gl/kdKNSs
- Bayarmagnai B, Nicolay BN, 1 Islam Abul BMMK, Lopez-Bigas N, Frolov MV (2012) Drosophila GAGA factor is required for full activation of the dE2f1-Yki/Sd transcriptional program. Cell Cycle 11: 4191-4202. Link: https://goo.gl/SvlLvs
- Dorogova NV, Fedorova EV, Bolobolova EU, Ogienko AA, Baricheva EM (2014) GAGA protein is essential for male germ cell development in Drosophila. Genesis 52: 738-751. Link: https://goo.gl/SqsZQt
- Kunwar PS, Starz-Gaiano M, Bainton RJ, Heberlein U, Lehmann R (2003) Tre1, a G protein-coupled receptor, directs transepithelial migration of Drosophila germ cells. PLoS Biol 1: e80. Link: https://goo.gl/CQq3Pe
- Kunwar PS, Sano H, Renault AD, Barbosa V, Fuse N, et al. (2008) Tre1 GPCR initiates germ cell transepithelial migration by regulating Drosophila melanogaster E-cadherin. J Cell Biol 183: 157-168. Link: https://goo.gl/1O72p5
- Sheng XR, Posenau T, Gumulak-Smith JJ, Matunis E, Van Doren M, et al. (2009) Jak–STAT regulation of male germline stem cell establishment during Drosophila embryogenesis. Dev Biol 334: 335-344. Link: https://goo.gl/sySfH6
- Li J, Xia F, Li WX (2003) Coactivation of STAT and Ras is required for germ cell proliferation and invasive migration in Drosophila. Dev Cell 5: 787-798. Link: https://goo.gl/mCNxOw
- Knipple DC, Seifert E, Rosenberg UB, Preiss A, Jäckle H (1985) Spatial and temporal patterns of Krüppel gene expression in early Drosophila embryos. Nature 317: 40. Link: https://goo.gl/3p43GM
- Rosenberg UB, Schröder C, Preiss A, Kienlin A, Côté S, et al. (1986) Structural homology of the product of the Drosophila Krüppel gene with Xenopus transcription factor IIIA . Nature 319: 336 – 339. Link: https://goo.gl/tyehA1
- Pankratz MJ, Hoch M, Seifert E, Jäckle H (1989) Krüppel requirement for knirps enhancement reflects overlapping gap gene activities in the Drosophila embryo . Nature 341: 337-340. Link: https://goo.gl/i7jUeB
- Štanojević D, Hoey T, Levine M (1989) Sequence-specific DNA-binding activities of the gap proteins encoded by hunchback and Krüppel in Drosophila. Nature 341: 331 – 335. Link:
- Treisman J, Desplan C (1989) The products of the Drosophila gap genes hunchback and Kriippel bind to the hunchback promoters. Nature (London) 341: 335-337. Link: https://goo.gl/aFwvYN
- Wieschaus E, Nüsslein-Volhard C, Jürgens G (1984) Mutations affecting the pattern of the larval cuticle inDrosophila melanogaster. Wilhelm Roux's archives of developmental biology 193: 296-307. Link: https://goo.gl/B6qehL
- Strecker TR, Merriam JR, Lengyel JA (1988) Graded requirement for the zygotic terminal gene, tailless, in the brain and tail region of the Drosophila embryo. Development 102: 721-734. Link: https://goo.gl/HTxnvP
- Casanova J (1990) Pattern formation under the control of the terminal system in the Drosophila embryo. Development 110: 621-628. Link: https://goo.gl/aJxH5U
- Pignoni F, Baldarelli RM, Steingrímsson E, Diaz RJ, Patapoutian A, et al. (1990) The Drosophila gene tailless is expressed at the embryonic termini and is a member of the steroid receptor superfamily. Cell 62: 151-163. Link: https://goo.gl/1TS2Nw
- Younossi-Hartenstein A, Green P, Liaw GJ, Rudolph K, Lengyel J, et al. (1997) Control of Early Neurogenesis of theDrosophilaBrain by the Head Gap Genestll, otd, ems, andbtd. Developmental biology 182: 270-283. Link: https://goo.gl/P46Aun
- Jiménez G, Guichet A, Ephrussi A, Casanova J (2000) Relief of gene repression by torso RTK signaling: role of capicua in Drosophila terminal and dorsoventral patterning. Gens Dev 14: 224-231. Link: https://goo.gl/BPSYok
- Paroush ZE, Wainwright SM, Ish-Horowicz D (1997) Torso signalling regulates terminal patterning in Drosophila by antagonising Groucho-mediated repression. Development 124: 3827-3834. Link: https://goo.gl/XcCqbz
- Chena YJ, Chiangb CS, Wengc LC, Lengyeld JA, Liawa GJ (2002) Tramtrack 69 is required for the early repression of tailless expression. Mechanisms of development 116: 75-83. Link: https://goo.gl/dm8wk8
- Ashe HL, Briscoe J (2006) The interpretation of morphogen gradients. Development 133: 385-394. Link: https://goo.gl/vUaJcW
- Lebrecht D, Foehr M, Smith E, Lopes FJ, Vanario-Alonso CE, et al. (2005) Bicoid cooperative DNA binding is critical for embryonic patterning in Drosophila. Proc Natl Acad Sci USA 102: 13176-13181. Link: https://goo.gl/gbEQFZ
- Liaw GJ, Rudolph KM, Huang JD, Dubnicoff T, Courey AJ, et al. (1995) The torso response element binds GAGA and NTF-1/Elf-1, and regulates tailless by relief of repression. Gens Dev 9: 3163-3176. Link: https://goo.gl/tyzTKn
- Staudt N, Fellert S, Chung HR, Jäckle H, Vorbrüggen G (2006) Mutations of the Drosophila zinc finger-encoding gene vielfältig impair mitotic cell divisions and cause improper chromosome segregation. Mol Biol Cell 17: 2356-2365. Link: https://goo.gl/dNPFnD
- Liang HL, Nien CY, Liu HY, Metzstein MM, Kirov N, et al. (2008) The zinc-finger protein Zelda is a key activator of the early zygotic genome in Drosophila. Nature 456: 400-403. Link: https://goo.gl/CAoG8v
- Yanez-Cuna JO, Dinh HQ, Kvon EZ, Shlyueva D, Stark A (2012) Uncovering cis-regulatory sequence requirements for context-specific transcription factor binding. Genome Research. 22: 2018-2030. Link: https://goo.gl/mx3WYr
- Foo SM, Sun Y, Lim B, Ziukaite R, O'Brien K, et al. (2014) Zelda potentiates morphogen activity by increasing chromatin accessibility. Curr Biol 24: 1341-1346. Link: https://goo.gl/RnU4ce
- Xu Z, Chen H, Ling J, Yu D, Struffi P, et al. Impacts of the ubiquitous factor Zelda on Bicoid-dependent DNA binding and transcription in Drosophila. Gens Dev 28: 608-621. Link: https://goo.gl/I4aQpf
- Harrison MM, Li XY, Kaplan T, Botchan MR, Eisen MB (2011) Zelda binding in the early Drosophila melanogaster embryo marks regions subsequently activated at the maternal-to-zygotic transition. PLoS Genet 7: e1002266. Link: https://goo.gl/Y2rqR0
- Satija R, Bradley RK (2012) The TAGteam motif facilitates binding of 21 sequence-specific transcription factors in the Drosophila embryo. Genome Res22: 656-665. Link: https://goo.gl/lrdw4F
- Li XY, Harrison MM, Villalta JE, Kaplan T, Eisen MB (2014) Establishment of regions of genomic activity during the Drosophila maternal to zygotic transition. Elife 3: e03737. Link: https://goo.gl/rlPH8C
- Bevilacqua A, Fiorenza MT, Mangia F (2000) A developmentally regulated GAGA box-binding factor and Sp1 are required for transcription of the hsp70. 1 gene at the onset of mouse zygotic genome activation. Development 127: 1541-1551. Link: https://goo.gl/AZvhS7
- Kim MH, Shin JS, Park S, Hur MW, Lee MO, et al. (2002) Retinoic acid response element in HOXA-7 regulatory region affects the rate, not the formation of anterior boundary expression. Int J Dev Biol 46: 325-328. Link: https://goo.gl/Tg5PXJ
- Kim SW, Ho SC, Hong SJ, Kim KM, So EC, et al. (2005) A novel mechanism of thyroid hormone-dependent negative regulation by thyroid hormone receptor, nuclear receptor corepressor (NCoR), and GAGA-binding factor on the rat cD44 promoter. J Biol Chem 280: 14545-14555. Link: https://goo.gl/Ckwcn9
- Min W, Woo HJ, Lee CJ, Lee KK, Yoon WK, et al. (1998) 307-bp Fragment in H0XA7 Upstream Sequence Is Sufficient for Anterior Boundary Formation. DNA and Cell Biology 17: 293-299. Link: https://goo.gl/BbZfCA
- Li J, Liang VCT, Sedgwick T, Wong J, Shi YB, et al. (1998) Unique organization and involvement of GAGA factors in transcriptional regulation of the Xenopus stromelysin-3 gene. Nucleic Acids Res 26: 3018-3025. Link: https://goo.gl/QqhbzA
- Galéra P, Musso M, Ducy P, Karsenty G (1994) c-Krox, a transcriptional regulator of type I collagen gene expression, is preferentially expressed in skin. Proc Natl Acad Sci USA 91: 9372-9376. Link: https://goo.gl/b77iwq
- Ghayor C, Herrouin JF, Chadjichristos C, Ala-Kokko L, Takigawa M, et al. (2000) Regulation of human COL2A1 gene expression in chondrocytes. Identification of C-Krox-responsive elements and modulation by phenotype alteration. J Biol Chem 275: 27421-27438. Link: https://goo.gl/pxNU4k
- Kypriotou M1, Beauchef G, Chadjichristos C, Widom R, Renard E, et al. (2007) Human collagen Krox up-regulates type I collagen expression in normal and scleroderma fibroblasts through interaction with Sp1 and Sp3 transcription factors. J Biol Chem 282: 32000-32014. Link: https://goo.gl/SPMGxk
- Galéra P, Park RW, Ducy P, Mattéi MG, Karsenty G (1996) c-Krox binds to several sites in the promoter of both mouse type I collagen genes. Structure/function study and developmental expression analysis. J Biol Chem 271: 21331-21339. Link: https://goo.gl/6t8ndp
- Vasanthi D, Anant M, Srivastava S, Mishra RK (2010) A functionally conserved boundary element from the mouse HoxD locus requires GAGA factor in Drosophila. Development 137: 4239-4247. Link: https://goo.gl/1JGTyF
Article Alerts
Subscribe to our articles alerts and stay tuned.
 This work is licensed under a Creative Commons Attribution 4.0 International License.
This work is licensed under a Creative Commons Attribution 4.0 International License.


 Save to Mendeley
Save to Mendeley
