1The First Hospital of Lanzhou University, Lanzhou City, Gansu Province 730000, China
2School of Basic Medical Science, Institute of Genetic, Lanzhou University, Lanzhou City, Gansu Province 730000, China
3The Secondary Hospital of Lanzhou University, Lanzhou City, Gansu Province 730000, China
4The Ningxia Hui Autonomous Region people’s Hospital, Yinchuan City, Ningxia Hui Autonomous Region 750000, China
Author and article information
Cite this as
Ji T, Tian C, Ji XZ, Huang L, Wang YC, et al. (2016) . . 2016; (): . Available from:
Copyright License
© 2016 Ji T, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.mtDNA: mitochondrial DNA; DNA: Deoxyribonucleic Acid; T2DM: Type 2 Diabetes Mellitus; PCR-RFLP: Polymerase Chain Reaction-the Restriction Fragment Length Polymorphism; ND3: NADH Dehydrogenase3; ROS: Reactive Oxygen Species; FPG: Fasting Plasma Glucose; ALT: Alanine Amino Transferase; TC: Total Cholesterol; TG: Triglyceride; BUN: Burine Nitrogen; SNP: Single Nucleotide Polymorphism;
Introduction
Diabetes mellitus type 2 (T2DM) is a long term metabolic disorder that is characterized by hyperglycemia, insulin resistance and relative lack of insulin[1]. Common symptoms include increased thirst, frequent urination, and unexplained weight loss. Long-term complications from hyperglycemia include heart disease, strokes, diabetic retinopathy which can result in blindness, kidney failure, and poor blood flow in the limbs which may lead to amputations [2-4]. Mitochondrial gene (mtDNA) mutation type diabetes mellitus has gradually attracted people’s attention[5]. Human mitochondrial DNA (mtDNA) is a double stranded circular molecule with 16569bp, which has a high mutation rate, and has become a heated focus on various diseases due to the lack of protection and efficient DNA repair mechanism[5-9]. The mtDNA oxidative phosphorylation system encompasses five (I–V) multi subunit enzyme complexes. Complex I consists of seven mitochondrial genomic encoded subunits (ND1, ND2, ND3,ND4L, ND4, ND5 and ND6) and 36 nuclear genes arrayed within the mitochondrial inner membrane[10]. The ND3 protein is a subunit of NADH dehydrogenase (ubiquinone), which is located in the mitochondrial inner membrane and is the largest of the five complexes of the electron transport chain. Position 10398 in the mtDNA is located in the gene encoding NADH dehydrogenase 3 (ND3). The G to A polymorphism in ND3 causes the substitution of alanine by threonine in amino acid 114 in the ND3 peptide. This amino acid substitution in the ND3 gene is hypothesized to affect protein function [11,12]. Darvishi et al. also reported that the G10398A polymorphism increased the risk of breast cancer in the Indian population [13]. In addition, mtDNA G10398A variant in African-American women with breast cancer provides resistance to apoptosis and promotes metastasis in mice [14]. Hui Xu also reported that Prognostic value of mitochondrial DNA content and G10398A polymorphism in non-small cell lung cancer [15]. At present, many investigations of association between T2DM and mtDNA from different races and territories found that frequencies of some mutations varied from areas and relevance with DM incidence, which meant that mutations may relate to DM morbidity or only be taken as natural polymorphism[16]. We aimed to test whether mitochondrial G10398A in ND3 locus, namely SNP (rs2853826), provided susceptibility to T2DM, from target population of Chinese northwestern T2DM patients, so as for new genetic clues of DM mechanism revealing.
Materials and Methods
Subjects
This study was approved by the Ethics Committee of Lanzhou University. After providing informed consent, 209 type 2 diabetic patients and 213 controls were recruited from the Hospital of Gansu Qingshui. All patients, who were Northwestern Chinese, were diagnosed according to the 1999 revised criteria for WHO; they did not have a family history of T2DM. The controls were age- and gender-matched unrelated healthy subjects recruited from individuals presenting for check-up in the hospital.
Genotyping analysis
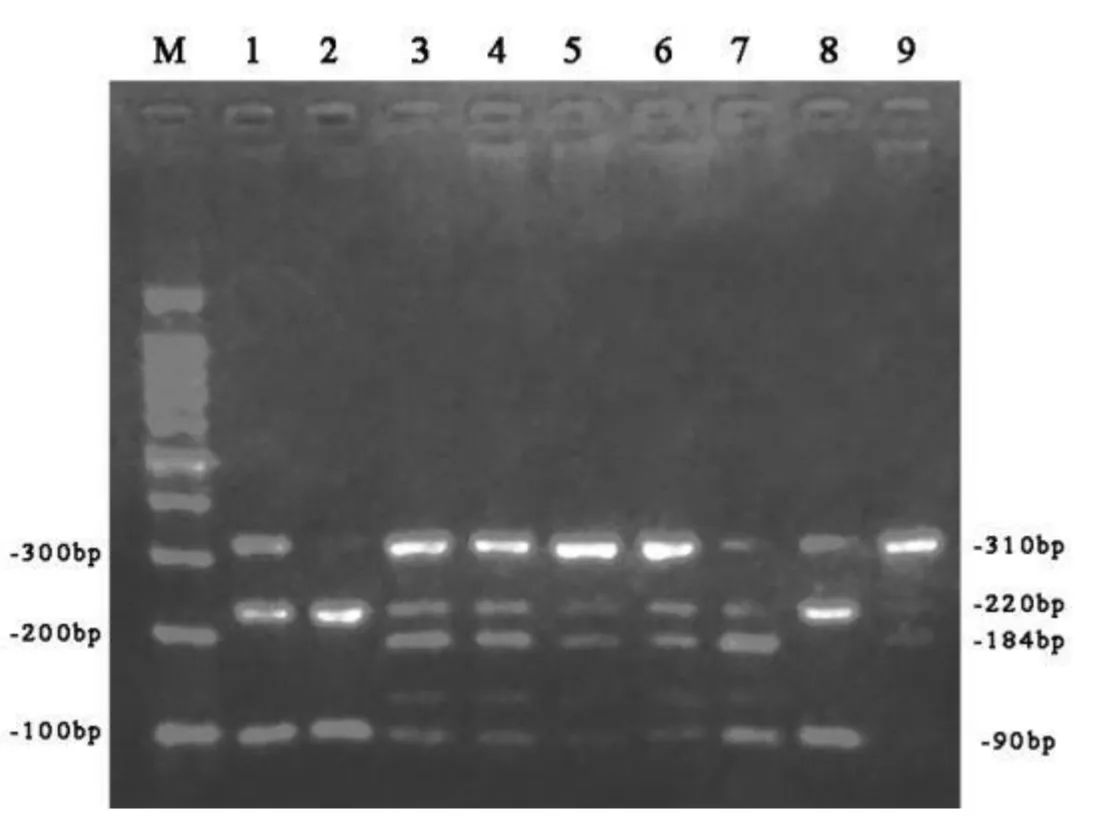
About 2 ml of venous blood was collected from each subject, using phenol chloroform method (Kunkel et al. 1977). The forward primer was 5´-tccttttacccctaccatgag-3´, and the reverse primer was 5´-tactatgcctagaaggaataat-3´. The PCR conditions were as follows. Initial denaturing was performed at 94 for 2min, and was followed by 30 cycles of denaturing at 94 for 30s, annealing at 54.3 for 30s, extension at 72for 45s, and final extension at 72 for 12 minutes. The PCR product was analyzed by electrophoresis in 2.0% agarose gel and visualized by ethidium bromide staining. The PCR product (310bp) was digested with DdeI (New England Biolabs, Beverly, MA) and separated on 2.5% agarose gel. The genotypic profiles were observed as three bands representing completely restricted fragments (sizes, 184bp, 90bp and 38bp) when G allele and two fragments 220bp and 90bp when an allele were present. Subsequent validation of the results was carried out through direct sequencing (Applied Biosystems, USA) to exclude any genotyping error or false positiveor negative results in PCR-RFLP.
Statistical analysis
Hardy–Weinberg equilibrium was assessed using χ2 test. Comparisons of genotype frequencies between T2DM patients and controls were performed using 2×2 contingency tables with χ2 analysis. Clinical characteristics were expressed as mean ± S.D. One-way analysis of variance was used to test for differences in means of phenotypic characteristics between genotypes. All P-values were two-tailed and P-values<0.05 were considered to be statistically significant. All statistical analyses were performed with SPSS for Windows (version 19.0IBM SPSS Statistics, USA).
Results
The genotypic profiles were observed as three bands representing completely restricted fragments (sizes, 184bp, 90bp and 38bp) when G allele and two fragments 220bp and 90bp when A allele were present (Figure 1).
In comparison with controls, cases had significantly higher levels of fasting plasma glucose (FPG), the plasma levels of total cholesterol(TC),triglyceride (TG), urea nitrogen (BUN)(P <0.05). However, there was no significance in age, the plasma levels of alanine transaminase (ALT) in both groups. This is consistent with diabetic features (Table 1).
The distributions of G10398A genotypes and alleles in theT2DM and control groups from a Chinese population are summarized in Table 2. The polymorphism (G10398A) fit well with the distributions expected under Hardy–Weinberg equilibrium for both groups. The genotype and allele distributions of the G10398A polymorphisms were not significantly different between theT2DM and control groups (Table 2). There is no association between the genotypes and clinical indicators T2DM group (Table 3).
Discussion
Type 2 diabetes mellitus (T2DM) affects a large population worldwide [17-19]. It was reported that the reactive oxygen species (ROS) overproduced by mitochondrion were associated with DM [20,21]. Mitochondrion is the main producer of ROS and its dysfunction causes superfluous ROS in insulin message pathway [22,23]. Meanwhile, the redundancy of ROS may accelerate cell apoptosis. It is widely accepted that high blood glucose and resultant oxidative stress mainly account for β cell functional injury, and elements, which affect ROS metabolism, can indirectly lead to the susceptibility to DM by influencing oxidative stress. Rare mtDNA mutation results in mitochondrion, β cell dysfunction and DM follows [24]. Bhat’s 2007 research demonstrated that mtDNAG10398A SNPs was missense mutation (amino acid change: Alanine → Threonine) and interfered β cell endocrine secretion by means of changing the conformation of the enzyme domain, decreasing the binding of transcription factors, preventing gene transcription and translation, debasing complexenzyme activity, less synthesis of ATP, excess of ROS [25-27]. This study suggested that G10398A mtDNA variance possibly played a role in providing susceptibility to T2DM in two North Indian populations.
We identified the genotypes of G10398A SNPs from Chinese northwestern population including 209 T2DM patients and 213 healthy controls. Frequencies of the genotypes and alleles were of no significant difference (P>0.05, χ2 test) and in no association with susceptibility to T2DM.In analyses of relationship between genotypes and clinic indicators, single transformation of genotypes has little influence on alternation of clinic indicators. Compared to the results of Bhat’s study in north India population, The probable reasons for the results were as followed: firstly, differentiated frequency distributions of mitochondrion gene mutations in various nations or populations; secondly, individual heterogeneity in mitochondrion gene mutation; additionally, diabetes is closely related to both the genetic and environmental elements, which have a close relationship with the individual life style and the environmental factors; moreover, the influence of sample size cannot be ignored.
Conclusion
To conclude, mtDNA G10398A variation has no association with T2MD in the present population group from the Chinese northwest. Therefore, further investigations are planned to include more SNPs or replicate in larger sample size to elucidate molecular mechanisms of predisposition caused by mtDNA mutation.
Article Alerts
Subscribe to our articles alerts and stay tuned.
 This work is licensed under a Creative Commons Attribution 4.0 International License.
This work is licensed under a Creative Commons Attribution 4.0 International License.

 Save to Mendeley
Save to Mendeley
