Journal of Clinical Microbiology and Biochemical Technology
Cord formation in Mycobacterium abscessus
Department of Microbiology, Faculty of Medicine, University of Colombo, Kynsey Road, Colombo 08, Sri Lanka
Author and article information
Cite this as
Adikaram C, Perera J, Perera GMM (2015) Cord formation in Mycobacterium abscessus. J Clin Microbiol Biochem Technol. 2015; 1(1): 18-19. Available from: 10.17352/jcmbt.000004
Copyright License
© 2015 Adikaram C, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.A sputum sample was received from a young woman with a past history of pulmonary mycobacterial disease (< 6 months). She has treated 6 month with first line anti tuberculosis drugs based on clinical symptoms and AFB microscopy (AFB positive) and she has cured. Next time, she came with cough and mild fever again. Molecular based laboratory identification of the sputum specimen confirmed that she has Mycobacterium abscessus infection.
After treating with sodium hydroxide (4%) (Sigma), it was centrifuged at 3000 g under refrigerated conditions (at 4°C). The centrifuged deposit was diluted in 1 ml of sterile distilled water to prepare the bacterial suspension. Two slopes of the Lowenstein–Jensen (L-J) medium (Difco), one containing paranitrobenzoic acid (PNB, Sigma) and 7H9 broth medium were inoculated with 100 μl of the bacterial suspension. A small portion of the bacterial suspension (~20 μl) was examined microscopically, using the Ziehl-Neelsen (ZN) stain, to determine the presence of acid fast bacilli (AFB). The inoculated media were incubated at 37°C in a 5% CO2 incubator. The phenotypic characters of colonies were observed and smears were prepared from cultures grown on all 3 media. Culture isolates were tested by nitrate reductase test.
Case Report
A sputum sample was received from a young woman with a past history of pulmonary mycobacterial disease (< 6 months). She has treated 6 month with first line anti tuberculosis drugs based on clinical symptoms and AFB microscopy (AFB positive) and she has cured. Next time, she came with cough and mild fever again. Molecular based laboratory identification of the sputum specimen confirmed that she has Mycobacterium abscessus infection.
After treating with sodium hydroxide (4%) (Sigma), it was centrifuged at 3000 g under refrigerated conditions (at 4°C). The centrifuged deposit was diluted in 1 ml of sterile distilled water to prepare the bacterial suspension. Two slopes of the Lowenstein–Jensen (L-J) medium (Difco), one containing paranitrobenzoic acid (PNB, Sigma) and 7H9 broth medium were inoculated with 100 µl of the bacterial suspension. A small portion of the bacterial suspension (~20 µl) was examined microscopically, using the Ziehl-Neelsen (ZN) stain, to determine the presence of acid fast bacilli (AFB). The inoculated media were incubated at 37°C in a 5% CO2 incubator. The phenotypic characters of colonies were observed and smears were prepared from cultures grown on all 3 media. Culture isolates were tested by nitrate reductase test.
The genomic DNA of bacterial culture was extracted by phenol chloroform method. The 240 bp fragment of the IS6110 insertion sequence (for identification of MTC members) [1] and 437 bp fragment of rpoB gene (for characterization of Mycobacterium species) (Kim et al. 2004) [2], were amplified by polymerase chain reaction (PCR) using extracted DNA. The specific primers and thermo-cycling parameters used for each amplification are shown in Table 1. The 50 µl PCR mixture containing 1.5 mM MgCl2, 200 µM of deoxynucleotide triphosphates (dNTPs) (Promega, USA), 1U Taq polymerase (GenScript), 20 pmol of each primer and 2.5 µl of genomic DNA (10 ng) was used for each PCR reaction. DNA extracted from H37Rv was used as the control.
The fragment of rpoB gene was PCR amplified in duplicate and both amplified products (40 µl, ~ 100 ng/µl) were custom DNA sequenced by Macrogen DNA sequencing service in Korea. A search of the GenBank database with the Basic Local Alignment Search Tool (BLAST) program was performed to determine the homology of the DNA sequences and homologues sequence were aligned with SeaView software (version 4.2.12) to identify the Mycobacterium species.
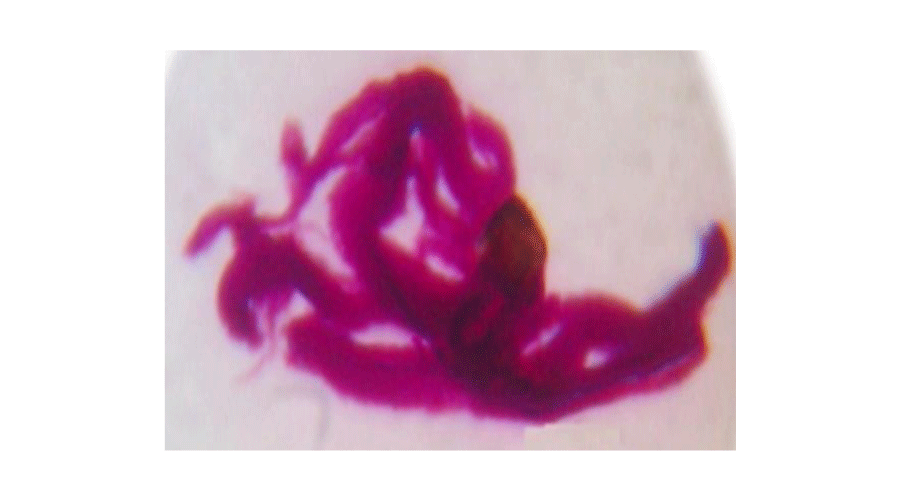
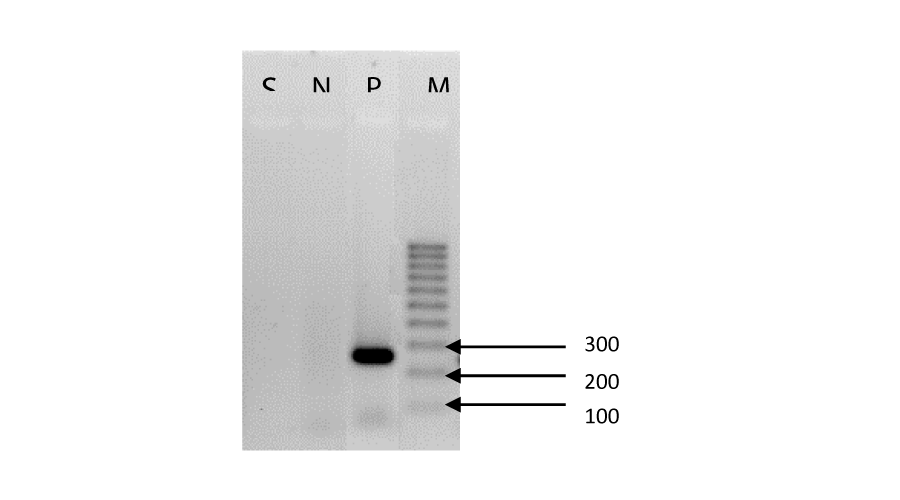
Rough cream coloured colonies after 6 days of incubation (Figure 1) were observed on both L-J and PNB incorporated L-J medium (Figure 1). The true cord was observed in the microscopic smear stained with Ziehl-Neelsen that was prepared from broth cultures (Figure 2). Both nitrate reductase test and DNA amplification of IS6110 fragment were negative (Figure 3). Rapid growth, positive growth in the presence of PNB and failure to PCR amplify the IS6110 fragment confirmed that the isolate is a non-tuberculosis Mycobacterium (NTM) species, although rough colony appearance and cord formation are characteristic features of MTC. The results of BLAST search and multiple alignment of the DNA sequence of fragment of rpoB gene confirmed that the isolate was Mycobacterium abscessus.
The analysis of DNA sequence of the rpoB gene is commonly used for the speciation of Mycobacterium genus [2,3]. Species level identification of NTM is medically important as their susceptibility to antibiotics vary with the species.
In some laboratories, the cord formation is considered as a distinctive feature of the MTC from NTM. Further, using microscopic observation drug susceptible test (MODS) which is used for detecting drug resistant tuberculosis is based on the presence of characteristics cording in drug containing culture medium [4]. However, the recent studies demonstrate that cord formation is not restricted to MCT and it is shared by few NTM species such as M. marinum and M. abscessus [5,6].
M. abscessus is an emerging pulmonary pathogen which shows a high rate of resistance for anti-tuberculosis drug regimens. Thus, infections caused by M. abscessus may be misdiagnosed as multidrug-resistant tuberculosis [6]. This is the first report of isolating M. abscessus from a clinical specimen in Sri Lanka and it clearly demonstrates that cord formation is not specific to MTC and detection of cording in broth culture should be further investigated before reaching a conclusion.
- Mulcahy GM, Kaminski ZC, Albanese EA, Sood R (1996 ) IS6110 based PCR methods for detection of Mycobacterium tuberculosis. J Clin Microbiol 34: 1348-1349
- Kim BJ, Hong SK, Lee KH (2004 ) Differential identification of Mycobacterium tuberculosis complex and non-tuberculosis mycobacteria by duplex PCR Assay using the RNA polymerase gene (rpoB). J Clin Microbiol 42: 1308-1312 .
- Kim BJ, Lee SH, Lyu MA (1999 ) Identification of mycobacterial species by comparative sequence analysis of the RNA polymerase gene (rpoB). J Clin Microbiol 37: 1714–1720 .
- Shiferaw G, Woldeamanuel Y, Gebeyehu M, Girmachew F, DemessieD, et al. (2007 ) Evaluation of Microscopic Observation Drug Susceptibility Assay for Detection of Multidrug-Resistant Mycobacterium tuberculosis. J Clin Microbiol 45: 1093–1097 .
- Staropoli JF, Branda JA (2008) Cord Formation in a Clinical Isolate of Mycobacterium marinum. J Clin Microbiol 46: 2814-2816.
- Sanchez-Chardi A, Olivares F, Byrd TF, Julian E, Brambilla C, et al. (2011 ) Demonstration of cord formation by Rough Mycobacterium abscessus variants: Implications for the Clinical Microbiology Laboratory. J Clin Microbiol 49: 2293–2295 .
Article Alerts
Subscribe to our articles alerts and stay tuned.
 This work is licensed under a Creative Commons Attribution 4.0 International License.
This work is licensed under a Creative Commons Attribution 4.0 International License.



 Save to Mendeley
Save to Mendeley
