Bonding Biology and Data: AI-Driven Advances in Stem Cell Research
Department of Biochemistry, Asutosh College (Affiliated to University of Calcutta), Kolkata, India
Author and article information
Cite this as
Karmakar K, et al. Bonding Biology and Data: AI-Driven Advances in Stem Cell Research. Ann Syst Biol . 2026; 9(1): 1-9. Available from: 10.17352/asb.000028
Copyright License
© 2026 Karmakar K, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.Stem cell–based therapies represent a central component of regenerative medicine owing to their capacities for self-renewal, multilineage differentiation, and disease modelling. However, their clinical translation remains hindered by biological heterogeneity, inconsistent differentiation outcomes, safety concerns, and limitations in scalability. Simultaneously, high-throughput experimental platforms have produced large, complex datasets that challenge conventional analytical methodologies. This review synthesises current literature addressing the application of artificial intelligence (AI), including machine learning and deep learning approaches, in stem cell research and therapy. Emphasis is placed on computational strategies for stem cell identification, characterisation, differentiation analysis, and cell fate prediction, drawing from studies integrating imaging and multi-omics data. Accumulating evidence indicates that AI-driven frameworks substantially enhance the accuracy, reproducibility, and efficiency of stem cell analyses. These approaches enable automated interpretation of high-dimensional datasets, facilitate prediction of lineage commitment, and improve quality assessment of cellular populations. AI methodologies further contribute to experimental optimisation and the development of predictive models supporting regenerative applications. Artificial intelligence is reshaping stem cell research by addressing longstanding analytical and biological challenges. Although technical, regulatory, and ethical limitations persist, continued advancements in AI integration are expected to accelerate the development of robust, scalable, and personalised regenerative therapies.
iPSCs: Induced Pluripotent Stem Cells; MSCs: Mesenchymal Stem Cells; ATMPs: Advanced Therapy Medicinal Products; CNN: Convolutional Neural Networks; GANs: Generative Adversarial Networks; OCT4: Octamer-binding transcription factor 4; NANOG: Nanog homeobox; SOX2: Sex-determining Region Y-box 2
Introduction
The growing complexity of stem cell biology characterized by vast datasets, heterogeneous cell populations, and intricate differentiation pathways, has made artificial intelligence an indispensable partner in advancing cell therapy from bench to bedside. Traditional empirical methods are increasingly inadequate for decoding the high-dimensional information generated by multi-omics, single-cell sequencing, and live imaging technologies. AI and machine learning now provide the analytical infrastructure required to interpret this complexity, enabling predictive modelling and process automation at unprecedented scales. In differentiation research, deep learning models analyse microscopy and molecular data to forecast optimal reprogramming conditions and assess cell identity non-invasively, dramatically accelerating protocol development. Virtual cell simulations, powered by generative networks and graph neural networks, further enhance this capability by predicting cellular responses to genetic or environmental perturbations with high fidelity. Quality control, a critical barrier to clinical translation, has been transformed through computer vision platforms that perform automated viability and purity assessments in real time, eliminating operator subjectivity and enabling scalable manufacturing. At the single-cell level, AI algorithms reconstruct developmental trajectories, identify rare but therapeutically relevant subpopulations, and predict functional outcomes such as immunomodulatory potential or tumorigenic risk insights that are essential for rational cell product selection. In the realm of personalised medicine, AI synthesises genomic, proteomic, and clinical data to forecast patient-specific responses, inform cell type selection, and guide dosing strategies, with demonstrated applications in hematopoietic transplantation and oncology. Manufacturing and biomaterials development have also benefited from reinforcement learning and algorithmic experimental design, which optimise parameters such as media formulation, oxygen tension, and shear stress to maximise yield and potency. Automated genome editing platforms further leverage AI to enhance editing efficiency and scalability. Together, these capabilities establish AI not merely as a convenience but as a core component of modern stem cell therapy, essential for translating biological complexity into safe, effective, and commercially viable regenerative treatments [1,2].
Types of artificial intelligence used in stem cell research
Artificial intelligence in stem cell research encompasses a diverse set of computational strategies designed to extract meaningful biological insight from complex, high-dimensional data. Rather than functioning as isolated tools, these approaches collectively enable pattern recognition, predictive modelling, and systems-level integration across molecular and phenotypic datasets. This section provides a conceptual overview of the principal AI paradigms employed in the field, while detailed applications are discussed in subsequent sections.
Machine learning
Machine learning (ML) refers to algorithms that learn statistical relationships directly from data to support classification, regression, and clustering tasks. In stem cell research, ML is widely applied to structured datasets such as gene expression profiles, proteomic measurements, and quantitative morphological descriptors. These models assist in identifying cell states, predicting lineage commitment, and uncovering hidden subpopulations within heterogeneous cultures. By modelling multidimensional biological relationships, ML provides scalable analytical capability beyond traditional rule-based or single-variable approaches [3].
Deep learning
Deep learning (DL), a specialised branch of machine learning, utilises multilayered neural networks to model complex and nonlinear biological systems. DL is particularly suited to unstructured data, including microscopy images and single-cell sequencing outputs, where it autonomously extracts hierarchical features without manual feature engineering [4]. In stem cell research, deep learning enables morphology-driven inference of pluripotency, differentiation dynamics, and cellular heterogeneity, thereby supporting real-time and non-invasive analytical workflows.
Generative artificial intelligence
Generative AI includes models capable of synthesising realistic biological data, thereby addressing challenges associated with limited sample availability. In stem cell studies, generative approaches support dataset augmentation, rare-cell representation, and simulation of potential biological states. These models enhance training robustness and open opportunities for computational hypothesis testing in silico before experimental validation [5].
AI for multi-modal integration
Stem cell systems are regulated across multiple biological layers, from genomic and epigenomic mechanisms to proteomic and metabolic networks. AI-driven integration frameworks learn shared representations across these modalities, enabling comprehensive mapping of cellular identity and functional state. Emerging interpretable AI models further aim to balance predictive performance with biological transparency, an important consideration for translational and regulatory contexts [6].
Beyond data analysis, AI increasingly contributes to experimental optimisation, quality assessment, and process standardisation in stem cell research. Predictive modelling supports protocol refinement, consistency monitoring, and safety evaluation, thereby bridging basic research and clinical translation.
Collectively, these AI paradigms form the computational foundation for the specialised applications discussed in Sections 3–5.
AI in stem cell identification and classification
Label-free pluripotency assessment and quality control
A transformative contribution of artificial intelligence to stem cell biology is the ability to assess pluripotency without recourse to destructive labelling techniques. Conventional approaches rely on immunocytochemical detection of transcription factors such as OCT4, NANOG, and SOX2, or on flow cytometric analysis of surface markers, both of which necessitate cell fixation or permeabilisation and irrevocably compromise subsequent experimental utility. Deep learning architectures trained on bright-field and phase-contrast microscopy images now enable the non-invasive prediction of pluripotency status directly from unmanipulated cellular morphology. These models exploit subtle, spatially distributed features invisible to the human eye—variations in nuclear texture, cytoplasmic granularity, colony edge definition, and intercellular packing density—that correlate reliably with molecular pluripotency networks [6]. The capacity to perform continuous, real-time monitoring of pluripotent stem cell cultures without perturbing the system represents a fundamental advance for regenerative medicine manufacturing, where batch-to-batch consistency, sterility maintenance, and product integrity are non-negotiable regulatory requirements. Furthermore, these label-free approaches permit longitudinal tracking of individual colonies throughout differentiation protocols, enabling the construction of high-resolution temporal maps of cell state transitions and the identification of intermediate populations that remain inaccessible to endpoint analytical methods [7].
Classification across stem cell compartments
The application of AI-based classification extends substantially beyond the pluripotent stem cell niche into specialised somatic stem cell compartments, each presenting unique analytical challenges. Mesenchymal stem cells, isolated from bone marrow, adipose tissue, and umbilical cord, exhibit pronounced morphological heterogeneity that correlates with donor age, passage number, and multilineage differentiation capacity. Convolutional neural networks trained on phase-contrast images of adherent MSC cultures successfully classify cells according to their osteogenic, adipogenic, or chondrogenic predisposition days before conventional lineage-specific markers become detectable. This predictive capacity carries considerable implications for autologous cell therapy manufacturing, where the functional potency of expanded cell products must be verified before patient administration [6,8,9].
Hematopoietic stem cell research has similarly benefited from AI-enabled image analysis. The functional distinction between long-term repopulating HSCs, short-term HSCs, and multipotent progenitors has historically required transplantation assays spanning weeks or months. Deep learning classifiers operating on high-resolution bright-field images now discriminate these subpopulations with accuracy approaching that of gold-standard immunophenotypic sorting. Critically, these morphological classifiers identify features—nuclear indentation profiles, mitochondrial distribution patterns, membrane ruffling dynamics—that correlate with quiescence depth and self-renewal capacity, offering mechanistic insights into the architectural basis of stem cell function that extend beyond mere sorting utility [8].
Cancer stem cell identification and therapeutic implications
Cancer stem cells represent a particularly formidable analytical target owing to their phenotypic plasticity, low abundance within heterogeneous tumour populations, and absence of universal, lineage-independent surface markers. Their morphological signatures remain poorly codified, with substantial variation observed across tissue of origin, oncogenic driver profile, and microenvironmental context. Deep learning models have demonstrated the capacity to detect putative CSC populations [10] from phase-contrast and fluorescence microscopy images of solid tumour cultures and haematological malignancies, identifying cells with distinct nuclear-to-cytoplasmic ratios, organelle distribution patterns, and substrate adhesion behaviours. In acute myeloid leukaemia, convolutional neural networks trained on imaging flow cytometry data successfully discriminate leukaemia stem cells from their normal haematopoietic counterparts—a distinction of profound clinical significance given that conventional chemotherapy frequently eradicates bulk blasts while sparing the chemotherapy-resistant stem cell reservoir responsible for relapse. The integration of generative adversarial networks to synthesise high-fidelity training images has partially addressed the chronic scarcity of expertly annotated CSC datasets, enabling model training on balanced class representations that substantially improve rare-cell detection sensitivity [11].
Methodological architectures and training paradigms
Contemporary deep learning applications in stem cell image analysis predominantly employ established convolutional neural network architectures originally developed for natural image classification tasks. DenseNet121, with its dense connectivity pattern permitting feature reuse across layers, demonstrates particular utility for stem cell applications where training datasets are modest in size [11]. MobileNetV2 provides a lightweight architecture amenable to deployment on standard laboratory workstation hardware without specialised graphics processing units. ResNet50’s skip connections facilitate training of substantially deeper networks without vanishing gradient phenomena, while VGG16, despite its relative architectural simplicity, remains widely employed as a feature extractor for transfer learning paradigms [6].
The training process typically commences with weights initialised from ImageNet-pretrained models, followed by fine-tuning on domain-specific stem cell image datasets. This transfer learning approach compensates for the relatively modest size of most publicly available stem cell image repositories and accelerates convergence. Data augmentation strategies—including random rotation, translation, scaling, elastic deformation, and intensity normalisation—further expand effective dataset size and improve model generalisation to unseen acquisition conditions. Emerging semi-supervised and weakly supervised paradigms reduce dependency on exhaustive pixel-level annotation, instead leveraging image-level labels or limited region annotations to guide learning. Iterative pseudo-balancing techniques specifically address the class imbalance problems endemic to stem cell imaging, where rare but biologically critical subpopulations may constitute less than one per cent of total cells [11].
Multimodal integration and emerging technological synergies
The contemporary trajectory of AI-enabled stem cell analysis increasingly emphasises integration across disparate data modalities, transcending the limitations inherent to image-only approaches. Alignment of morphological features extracted from microscopy data with transcriptional profiles derived from single-cell RNA sequencing enables the construction of multimodal cell atlases wherein physical appearance is quantitatively linked to gene regulatory network activity [12]. Deep learning frameworks analysing training dynamics across annotated single-cell datasets identify mislabelled cells, resolve transcriptional states existing along continuous differentiation trajectories, and infer pseudo-temporal ordering from binary labels alone—tasks that previously required bespoke trajectory inference algorithms or extensive prior knowledge of lineage relationships [13].
Simultaneously, the convergence of artificial intelligence with organoid technology has generated powerful platforms for developmental biology and drug discovery. Organoids recapitulate key aspects of human tissue architecture and function, but generate extraordinarily complex imaging datasets prohibitive to comprehensive manual analysis. AI systems automate the quantification of organoid number, size, morphology, budding frequency, and interior cyst formation across thousands of structures per experimental condition. These systems detect treatment effects and genetic perturbation phenotypes with sensitivity exceeding human observation while operating at throughput scales compatible with primary small molecule screening campaigns [10].
AI-driven stem cell differentiation and fate prediction
Biological complexity of differentiation processes
Stem cell differentiation is governed by a complex interplay of genetic, epigenetic, and environmental factors. Lineage commitment is not a linear or deterministic process; rather, it involves dynamic transitions through intermediate cellular states influenced by signalling pathways, transcriptional networks, and microenvironmental cues. Cellular heterogeneity and stochastic fluctuations further complicate the prediction of differentiation outcomes [3].
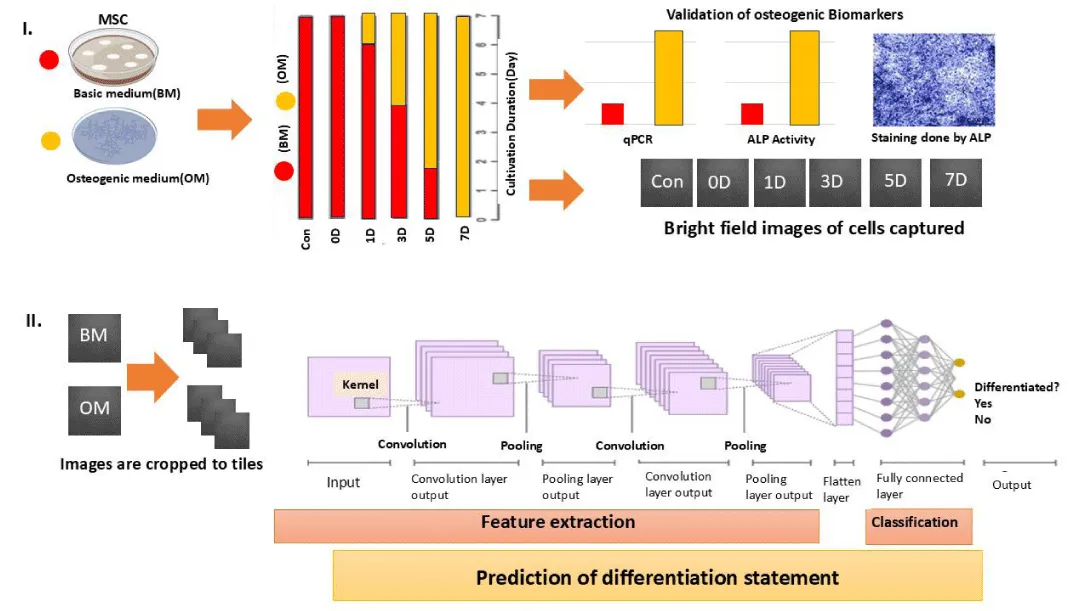
Traditional analytical approaches often rely on static snapshots of gene expression or marker profiles, limiting their ability to capture temporal dynamics and nonlinear dependencies. Consequently, there is a growing recognition that predictive modelling of differentiation requires computational frameworks capable of learning complex, multivariate relationships. The integration of morphology-derived imaging features with osteogenic induction dynamics and computational classification strategies is conceptually illustrated in Figure 1.
Machine learning approaches to lineage prediction
Machine learning models have been applied to predict stem cell fate based on molecular and phenotypic features. Early studies demonstrated that supervised learning algorithms could classify differentiation outcomes using transcriptomic signatures, providing proof-of-concept for computational fate prediction. These approaches highlighted the potential of ML to identify key regulatory features associated with lineage commitment [1,14].
More recent studies have expanded upon this foundation by incorporating larger datasets and more sophisticated models. Feature selection techniques enable the identification of genes and pathways most strongly associated with specific differentiation trajectories, offering mechanistic insight alongside predictive power [1].
Deep learning and single-cell resolution
The integration of deep learning with single-cell technologies represents a major advance in stem cell research. Single-cell RNA sequencing captures transcriptional heterogeneity at unprecedented resolution but generates sparse and noisy data. Deep generative models, such as variational autoencoders, address these challenges by learning low-dimensional representations that preserve biologically meaningful variation [11].
These models facilitate the reconstruction of differentiation trajectories and enable the identification of transient intermediate states that are difficult to capture experimentally. Importantly, deep learning frameworks can model temporal dynamics, allowing prediction of future cellular states based on early transcriptional patterns.
Recent AI approaches have also been used to predict cellular responses to genetic or chemical perturbations, thereby enabling in silico exploration of differentiation strategies. Such predictive capabilities reduce experimental burden and accelerate protocol optimization [5].
AI-guided optimisation of differentiation protocols
Beyond analysis, AI is increasingly employed as a decision-support tool for optimising differentiation protocols. By integrating experimental data across multiple conditions, machine learning models can identify optimal combinations of growth factors, signalling molecules, and culture parameters. This reduces reliance on empirical trial-and-error approaches and enhances reproducibility across laboratories.
From a translational perspective, AI-guided differentiation offers a pathway toward standardised and scalable production of therapeutic cell types. Predictive modelling ensures consistent lineage specification while minimising off-target differentiation, thereby improving safety and efficacy [12].
Integration of Artificial intelligence with single-cell and multi-omics technologies
Emergence of high-dimensional omics data in stem cell research
Recent advances in high-throughput sequencing and mass spectrometry technologies have enabled comprehensive molecular profiling of stem cells at unprecedented resolution. Single-cell RNA sequencing (scRNA-seq), chromatin accessibility assays (scATAC-seq), proteomics, and metabolomics collectively capture multiple layers of cellular regulation. These technologies have been instrumental in uncovering transcriptional heterogeneity, lineage bifurcations, and rare cell states that are often obscured in bulk analyses [14].
However, the integration and interpretation of such high-dimensional datasets present significant analytical challenges. Stem cell datasets are often sparse, noisy, and affected by batch effects arising from experimental variability. Conventional statistical methods struggle to integrate heterogeneous data types while preserving biologically meaningful signals. Consequently, there is a growing reliance on AI-based frameworks to manage the scale and complexity of multi-omics stem cell data [4,11,13].
Role of AI in multi-omics data integration
Artificial intelligence plays a central role in integrating diverse omics datasets by learning shared latent representations across multiple data modalities. Machine learning approaches such as matrix factorisation, manifold learning, and deep generative models enable dimensionality reduction while retaining critical biological information. These models facilitate the identification of regulatory programs that govern stem cell maintenance, differentiation, and lineage commitment [15].
Multi-omics integration using AI allows the simultaneous analysis of transcriptional, epigenetic, and proteomic features, offering a systems-level understanding of stem cell behaviour. Such integrative analyses are particularly valuable for identifying molecular signatures associated with pluripotency, differentiation potential, and cellular stability. Importantly, AI-driven integration reduces human bias by uncovering data-driven relationships rather than relying solely on predefined biological assumptions [14,15].
Biomarker discovery for stem cell potency and safety
Ensuring the safety and efficacy of stem cell-based therapies requires robust biomarkers that reliably indicate cell potency, differentiation status, and genetic stability. AI-based analysis of multi-omics data enables the identification of predictive biomarkers by correlating molecular features with functional outcomes. These biomarkers can be used to assess tumorigenic potential, differentiation efficiency, and long-term stability of stem cell-derived products [2].
By learning complex nonlinear associations between molecular features and phenotypic outcomes, AI models enhance the sensitivity and specificity of biomarker discovery. This is particularly critical for clinical applications, where early detection of aberrant cell states can prevent adverse outcomes following transplantation [16].
Patient-derived iPSCs and personalised regenerative medicine
The convergence of AI and patient-derived induced pluripotent stem cells has opened new avenues for personalised regenerative medicine. IPSCs generated from individual patients retain disease-specific genetic and epigenetic signatures, enabling personalised disease modelling and therapeutic testing. AI-driven analysis of patient-specific datasets allows prediction of disease progression, drug responses, and therapeutic efficacy [5,17]. Such predictive frameworks support the development of individualised treatment strategies by tailoring stem cell-based interventions to patient-specific molecular profiles. In this context, AI serves as a bridge between experimental data and clinical decision-making, facilitating the translation of personalised regenerative therapies.
Challenges in single-cell and multi-omics integration
Despite its promise, AI-driven omics integration faces several limitations. Data sparsity, technical noise, and limited availability of well-annotated datasets can compromise model performance. Additionally, the interpretability of complex AI models remains a challenge, particularly when integrating multiple data modalities. Addressing these issues requires standardised data generation protocols, transparent model architectures, and close collaboration between computational scientists and experimental biologists [4].
AI in drug screening and disease modelling using stem cells
Patient-specific iPSCs and AI-assisted analysis
Concomitantly, AI-assisted phenotyping of patient-derived iPSCs has emerged as a key application of AI technology in disease modelling and drug screening. AI-assisted image analysis can facilitate the quantitative assessment of cellular phenotypes, allowing for the identification of subtle changes that may not be detectable by the naked eye. For instance, the contractility of patient-specific iPSC-derived cardiomyocytes can be evaluated by AI-assisted image analysis for early screening of cardiotoxic drug responses. In addition, ML-based analysis can be employed to classify disease-specific morphological phenotypes of iPSC-derived motor neurons from patients with neurodegenerative diseases such as amyotrophic lateral sclerosis (ALS). Such AI-assisted phenotyping can provide not only improved predictive outcomes for patient-specific drug responses but also diminished experimental errors in iPSC studies [17,18]. In a recent report, more than 85% of disease-specific phenotypes could be successfully identified by AI-assisted phenotyping, suggesting that AI technology has the potential to provide valuable information for the identification of disease-specific signatures in patient-specific iPSCs [19,20]. Representative examples of such AI-enabled phenotypic classification strategies across diverse stem cell–derived models are summarised in Table 1.
Organoids in modelling complex diseases
Aside from single cell types, AI-assisted analysis has also been applied to 3D models such as organoids. Organoids are 3D tissues generated from iPSCs that more accurately represent the structure and cellular diversity of human tissues compared to conventional 2D cultured cells. AI-assisted analysis of high-content imaging of organoids can monitor the differentiation process, assess cellular diversity, and predict the functional response to drugs. Organoid-based approaches to disease modelling, drug screening, and toxicity testing are enabled by AI-assisted analysis and provide scalable and high-throughput platforms for drug development [18]. For example, AI-assisted analysis can identify aberrant morphological features of organoids that may not yet display apparent disease phenotypes, and may thus allow for earlier intervention. Such approaches expedite drug development and improve the predictive capacity of in vitro models for personalised regenerative medicine [19].
High-throughput drug screening and predictive modelling
AI-assisted screening approaches can incorporate imaging, transcriptomic, and proteomic data to enable high-throughput drug discovery. It can predict cellular responses to candidate drugs, off-target effects, and dosing regimens before screening. These methods can be faster and more cost-effective than traditional screening methods and offer a better understanding of the drugs’ mechanisms. Recent reports suggest that AI-driven screening frameworks can substantially reduce experimental burden and candidate attrition rates. This predictive ability is particularly important for stem cell-based treatments, which need to be effective and safe across patient-specific cells [21].
Consequences and future directions
The integration of AI with stem cell technology has the potential to enhance disease modelling and drug screening by learning and predicting complex cellular responses to improve the efficiency of candidate drug identification and patient stratification. With the development of more interpretable AI models and their integration with automated culture platforms, the future of AI in regenerative medicine and precision therapeutics may include the ability to predict cellular and patient-level responses to candidate drugs, bringing precision therapeutics to the forefront of the field. A continued push to develop robust, interpretable AI models will be essential to realising the potential of these technologies in regenerative medicine and precision therapeutics [11,12].
AI technologies enabling quality control
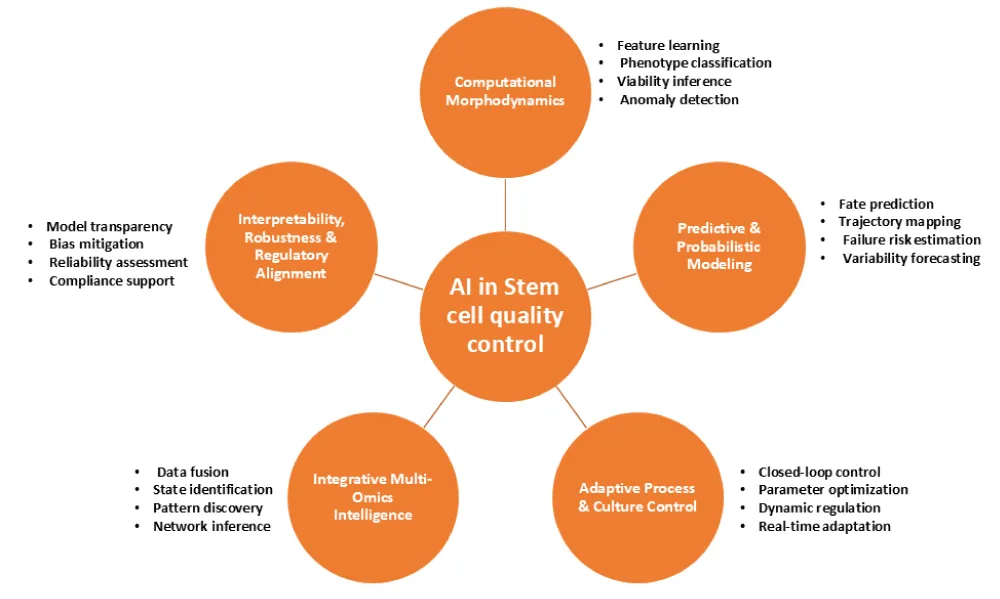
AI has brought transformative capabilities to the quality control of stem cell cultures, moving beyond simple replacement of manual observation to enable automated, real-time, and predictive monitoring of critical culture attributes. By integrating advanced computational methods, AI can continuously assess cell morphology, colony integrity, and other essential characteristics, providing insights that are difficult or impossible to capture through conventional microscopy. Figure 2 illustrates how different AI techniques correspond to specific quality-monitoring challenges, highlighting the rationale for selecting tools for particular critical quality attributes (CQAs). Traditional microscopy depends heavily on manual interpretation, which is time-consuming, subjective, and limited in scalability. In contrast, AI-driven machine vision, especially convolutional neural networks (CNNs), facilitates high-throughput and objective analysis of stem cell images. CNN classifiers have demonstrated high accuracy in detecting morphological markers of pluripotency, often exceeding human-level reproducibility while enabling scalable, objective assessment. These models are sensitive enough to detect subtle morphological variations, such as changes in colony shape, granularity, or density, that often escape human observation [2].
Ethical and regulatory considerations in AI-integrated stem cell research
While artificial intelligence has introduced transformative analytical capacity into stem cell research, its integration into experimental and translational workflows raises ethical and regulatory questions that extend beyond general healthcare AI concerns. In the context of stem cell–based therapies, algorithmic decisions may directly influence assessments of cell potency, differentiation status, genomic stability, and tumorigenic risk. Therefore, the reliability and interpretability of AI systems become matters of clinical safety rather than merely computational performance. One critical consideration is algorithmic transparency. Many advanced models, particularly deep learning systems, operate as high-dimensional predictive frameworks whose internal decision pathways are not easily interpretable. When such models are used to evaluate pluripotency markers or predict malignant transformation risk, regulatory agencies are likely to require evidence of biological rationale, reproducibility across independent datasets, and robustness against batch effects or culture variability. Black-box predictions without mechanistic insight may face barriers to clinical approval, particularly in applications involving human implantation.
Data provenance and standardisation represent additional challenges. AI systems trained on stem cell imaging, multi-omics data, or differentiation assays may inherit biases arising from laboratory-specific protocols, donor variability, or imaging platforms. Without harmonised data acquisition standards, predictive models risk reduced generalizability across institutions. Regulatory evaluation frameworks will therefore need to consider not only algorithmic accuracy but also dataset representativeness and cross-site validation. A particularly sensitive issue concerns tumorigenicity prediction. Pluripotent stem cells inherently possess proliferative potential, and incomplete differentiation or residual undifferentiated cells may pose oncogenic risk. If AI algorithms are employed to predict tumorigenic propensity based on molecular or morphological signatures, questions arise regarding validation thresholds, acceptable false-negative rates, and long-term monitoring responsibilities. Establishing clear performance benchmarks will be essential before such systems can inform clinical decision-making [2,18].
In addition, the use of patient-derived induced pluripotent stem cells introduces ethical considerations related to consent and secondary data usage. AI-driven analyses often require large-scale data aggregation, including genomic and phenotypic information. Transparent governance policies must ensure that donor consent extends to computational modelling applications and that privacy safeguards are maintained when datasets are shared for algorithm training [16].
Looking forward, regulatory agencies may increasingly require co-development pathways in which AI tools and stem cell products are evaluated together rather than independently. This integrated assessment approach would recognise that algorithmic outputs may influence release criteria, manufacturing consistency, and safety profiling. Collaborative dialogue between computational scientists, stem cell biologists, clinicians, and regulatory authorities will therefore be essential to establish standardised validation protocols. Ultimately, ethical deployment of AI in stem cell research depends not only on technological sophistication but also on accountability, transparency, and rigorous validation [2]. Ensuring that computational predictions align with biological reality and clinical safety standards will be fundamental to translating AI-assisted regenerative strategies into responsible therapeutic practice.
Future perspectives: AI-driven personalised regenerative medicine
The convergence of artificial intelligence (AI) and stem cell technologies is poised to redefine personalised regenerative medicine. While traditional precision medicine has focused on integrating genomic, transcriptomic, and clinical data to tailor treatments, the next decade is likely to see AI enabling patient-specific stem cell therapies that are both predictive and adaptive. Machine learning models can analyse multi-omics datasets, cellular imaging, and patient clinical histories to forecast how individual patients’ cells will respond to specific therapies, guiding the selection of optimal cell types, differentiation protocols, and therapeutic dosing [1,15].
A critical frontier lies in scaling stem cell production for clinical applications. AI-driven bioprocess optimisation can model and adjust culture conditions in real time, ensuring consistent cell quality and potency while minimising variability. Similarly, predictive algorithms can help anticipate regulatory hurdles and clinical adoption timelines, offering insights into safety, efficacy, and long-term outcomes. By simulating patient-specific responses in silico, AI has the potential to reduce experimental burden, accelerate preclinical development, and support more informed decision-making for both clinicians and regulatory bodies.
Beyond efficiency, AI may enhance disease-specific personalisation. For example, patient-derived iPSCs and organoids can be paired with AI to model individual disease progression, predict therapeutic responses, and identify the most effective interventions. This capability could be particularly transformative for complex or rare conditions, where traditional clinical data are limited. Moreover, integrating explainable AI techniques ensures that predictions remain interpretable and actionable, fostering clinician trust and regulatory acceptance.
Looking forward, AI-driven regenerative medicine promises a future where therapies are precisely matched to patient biology, not only improving efficacy but also reducing adverse effects. As computational models become increasingly robust, their integration with automated cell culture systems, high-content imaging, and real-time monitoring will allow the development of adaptive, self-optimising treatment strategies, effectively bridging the gap between laboratory innovation and clinical application [11].
Conclusion
Artificial intelligence is transforming stem cell therapy by addressing fundamental challenges in differentiation, characterisation, and scalability. Through predictive modelling, automation, and integrative analysis, AI enhances reproducibility, efficiency, and translational potential. While significant challenges remain, continued integration of AI with stem cell research promises to accelerate the development of safe, effective, and personalised regenerative therapies. The future of stem cell therapy will increasingly depend on the synergistic collaboration between biological insight and computational intelligence.
Conflict of interest
The authors declared that this work was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgement
The authors would like to acknowledge the Research and Development Cell, Asutosh College, Kolkata, West Bengal, for their consistent support and for providing access to the necessary resources and literature. Figures in this review were created with BioRender. The authors contributed equally to all aspects of this work.
- Ali M, Benfante V, Basirinia G, Alongi P, Sperandeo A, Quattrocchi A, et al. Applications of artificial intelligence, deep learning, and machine learning to support the analysis of microscopic images of cells and tissues. J Imaging. 2025;11(2):59. Available from: https://doi.org/10.3390/jimaging11020059
- Leonov GE, Grinchevskaya LR, Makhnach OV, Samburova MV, Goldshtein DV, Salikhova DI. Safety assessment of stem cell-based therapies: Current standards and advancing frameworks. Cells. 2025;14(21):1660. Available from: https://doi.org/10.3390/cells14211660
- Choudhery MS, Arif T, Mahmood R. Applications of artificial intelligence in stem cell therapy. World J Stem Cells. 2025;17(8). Available from: https://doi.org/10.4252/wjsc.v17.i8.106086
- Huang M, Dissanayaka WL, Yiu CKY. Artificial intelligence driven innovation: Advancing mesenchymal stem cell therapies and intelligent biomaterials for regenerative medicine. Bioengineering. 2025;12(12):1302. Available from: https://doi.org/10.3390/bioengineering12121302
- Marzec-Schmidt K, Ghosheh N, Stahlschmidt SR, Küppers-Munther B, Synnergren J, Ulfenborg B. Artificial intelligence supports automated characterization of differentiated human pluripotent stem cells. Stem Cells. 2023;41(9):850-861. Available from: https://doi.org/10.1093/stmcls/sxad049
- Priyaka IGN K. Deep learning: Automating the future of stem cell bioimage analysis. Int J Sci Adv. 2024;5(6). Available from: https://doi.org/10.51542/ijscia.v5i6.59
- Li P, Li Y, Wang C. The paradigm shift in neural stem cells basic research driven by artificial intelligence related technologies. Front Cell Neurosci. 2025;19. Available from: https://doi.org/10.3389/fncel.2025.1696943
- Akiyoshi R, Hase T, Sathiyananthavel M, Ghosh S, Kitano H, Yachie A. Noninvasive, label-free image approaches to predict multimodal molecular markers in pluripotency assessment. Sci Rep. 2024;14(1). Available from: https://doi.org/10.1038/s41598-024-66591-z
- Shi Q, Song F, Zhou X, Chen X, Cao J, Na J, et al. Early predicting osteogenic differentiation of mesenchymal stem cells based on deep learning within one day. Ann Biomed Eng. 2024;52(6):1706-1718. Available from: https://doi.org/10.1007/s10439-024-03483-3
- Hwang WL, Huang SW, Hsiao AC, Chen CY, Hsu KF, Hsieh YT, et al. Harnessing engineered mesenchymal stem cell-derived extracellular vesicles for innovative cancer treatments. Stem Cell Res Ther. 2025;16:648. Available from: https://doi.org/10.1186/s13287-025-04708-5
- Solopov M, Chechekhina E, Turchin V, Popandopulo A, Filimonov D, Burtseva A, et al. Current trends and future opportunities of AI-based analysis in mesenchymal stem cell imaging: A scoping review. J Imaging. 2025;11(10):371. Available from: https://doi.org/10.3390/jimaging11100371
- Soori M, Jough FKG, Dastres R, Arezoo B. AI-based decision support systems in Industry 4.0, a review. J Econ Technol. 2025;4:206-225. Available from: https://doi.org/10.1016/j.ject.2024.08.005
- Choudhury M, Deans AJ, Candland DR, Deans TL. Advancing cell therapies with artificial intelligence and synthetic biology. Curr Opin Biomed Eng. 2025;34:100580. Available from: https://doi.org/10.1016/j.cobme.2025.100580
- Lim J, Park C, Kim M, Kim H, Kim J, Lee D. Advances in single-cell omics and multiomics for high-resolution molecular profiling. Exp Mol Med. 2024;56(3):515-526. Available from: https://doi.org/10.1038/s12276-024-01186-2
- Ballard JL, Wang Z, Li W, Shen L, Long Q. Deep learning-based approaches for multi-omics data integration and analysis. Iodate Min. 2024;17:38. Available from: https://doi.org/10.1186/s13040-024-00391-z
- Mennella C, Maniscalco U, De Pietro G, Esposito M. Ethical and regulatory challenges of AI technologies in healthcare: A narrative review. Heliyon. 2024;10(4):e26297. Available from: https://doi.org/10.1016/j.heliyon.2024.e26297
- Johnson KB, Wei Q, Weeraratne D, Frisse ME, Misulis K, Rhee K, et al. Precision medicine, AI, and the future of personalized health care. Clin Transl Sci. 2020;14(1):86-93. Available from: https://doi.org/10.1111/cts.12884
- Kusumoto D, Yuasa S, Fukuda K. Induced pluripotent stem cell-based drug screening by use of artificial intelligence. Pharmaceuticals. 2022;15(5):562. Available from: https://doi.org/10.3390/ph15050562
- Chapman T, Lassmann T. Single-cell data combined with phenotypes improves variant interpretation. BMC Genomics. 2025;26:540. Available from: https://doi.org/10.1186/s12864-025-11711-w
- Du X, Chen Z, Li Q, Yang S, Jiang L, Yang Y, et al. Organoids revealed: Morphological analysis of the profound next-generation in-vitro model with artificial intelligence. Bio-des Manuf. 2023;6:319-339. Available from: https://doi.org/10.1007/s42242-022-00226-y
- Chen X, Xu H, Yu S, Hu W, Zhang Z, Wang X, et al. AI-driven transcriptome prediction in human pathology: From molecular insights to clinical applications. Biology. 2025;14:651. Available from: https://doi.org/10.3390/biology14060651
Article Alerts
Subscribe to our articles alerts and stay tuned.
 This work is licensed under a Creative Commons Attribution 4.0 International License.
This work is licensed under a Creative Commons Attribution 4.0 International License.


 Save to Mendeley
Save to Mendeley
